from sklearn import datasets
from sklearn.neighbors import KNeighborsClassifier
from sklearn.model_selection import (train_test_split,
cross_val_score,
GridSearchCV,
RandomizedSearchCV)
from sklearn.linear_model import (LinearRegression,
Ridge,
Lasso,
LogisticRegression,
ElasticNet)
from sklearn.metrics import (mean_squared_error,
classification_report,
confusion_matrix,
roc_curve,
roc_auc_score,
precision_recall_curve,
plot_precision_recall_curve)
from sklearn.tree import DecisionTreeClassifier
from sklearn.impute import SimpleImputer
from sklearn.pipeline import Pipeline
from sklearn.svm import SVC # Support Vector Classiffication
from sklearn.preprocessing import (scale, StandardScaler)
import pandas as pd
import numpy as np
from scipy.stats import randint
import seaborn as sns
import matplotlib.pyplot as plt
%matplotlib inline
import warnings
warnings.filterwarnings("ignore")
plt.style.use("ggplot")Overview
Machine learning is the field that teaches machines and computers to learn from existing data to make predictions on new data: Will a tumor be benign or malignant? Which of your customers will take their business elsewhere? Is a particular email spam? We will use Python to perform supervised learning, an essential component of machine learning. We will build predictive models, tune their parameters, and determine how well they will perform with unseen data—all while using real world datasets. We be using scikit-learn, one of the most popular and user-friendly machine learning libraries for Python.
Libraries
Classification
Introduction to classification problems and how to solve them using supervised learning techniques. We’ll explore a political dataset, where we will classify the party affiliation of United States congressmen based on their voting records.
Supervised learning
What is machine learning?
- The art and science of:
- Giving computers the ability to learn to make decisions from data
- without being explicitly programmed!
- Examples:
- Learning to predict whether an email is spam or not
- Clustering wikipedia entries into different categories
- Supervised learning: Uses labeled data
- Unsupervised learning: Uses unlabeled data
Unsupervised learning
- Uncovering hidden patterns from unlabeled data
- Example:
- Grouping customers into distinct categories (Clustering)
Reinforcement learning
- Software agents interact with an environment
- Learn how to optimize their behavior
- Given a system of rewards and punishments
- Draws inspiration from behavioral psychology
- Applications
- Economics
- Genetics
- Game playing
- AlphaGo: First computer to defeat the world champion in Go
Supervised learning
- Predictor variables/features and a target variable
- Aim:
- Predict the target variable, given the predictor variables
- Classication: Target variable consists of categories
- Regression: Target variable is continuous
Naming conventions
- Features = predictor variables = independent variables
- Target variable = dependent variable = response variable
Supervised learning
- Automate time-consuming or expensive manual tasks
- Example: Doctor’s diagnosis
- Make predictions about the future
- Example: Will a customer click on an ad or not?
- Need labeled data
- Historical data with labels
- Experiments to get labeled data
- Crowd-sourcing labeled data
Supervised learning in Python
- We will use scikit-learn/sklearn
- Integrates well with the SciPy stack
- Otherlibraries
- Tensor Flow
- keras
Exploratory data analysis
The Iris dataset
- Features:
- Petal length
- Petal width
- Sepal length
- Sepal width
- Target variable:
- Species Versicolor
- Virginica
- Setosa
The Iris dataset in scikit-learn
iris = datasets.load_iris()
type(iris)sklearn.utils.Bunchiris.keys()dict_keys(['data', 'target', 'frame', 'target_names', 'DESCR', 'feature_names', 'filename'])type(iris.data)numpy.ndarraytype(iris.target)numpy.ndarrayiris.data.shape(150, 4)iris.target_namesarray(['setosa', 'versicolor', 'virginica'], dtype='<U10')Exploratory data analysis (EDA)
X = iris.data
y= iris.target
df = pd.DataFrame(X, columns=iris.feature_names)
df.head()| sepal length (cm) | sepal width (cm) | petal length (cm) | petal width (cm) | |
|---|---|---|---|---|
| 0 | 5.1 | 3.5 | 1.4 | 0.2 |
| 1 | 4.9 | 3.0 | 1.4 | 0.2 |
| 2 | 4.7 | 3.2 | 1.3 | 0.2 |
| 3 | 4.6 | 3.1 | 1.5 | 0.2 |
| 4 | 5.0 | 3.6 | 1.4 | 0.2 |
df2 = df.copy()
df2['target_names'] = iris.target
df2.head()| sepal length (cm) | sepal width (cm) | petal length (cm) | petal width (cm) | target_names | |
|---|---|---|---|---|---|
| 0 | 5.1 | 3.5 | 1.4 | 0.2 | 0 |
| 1 | 4.9 | 3.0 | 1.4 | 0.2 | 0 |
| 2 | 4.7 | 3.2 | 1.3 | 0.2 | 0 |
| 3 | 4.6 | 3.1 | 1.5 | 0.2 | 0 |
| 4 | 5.0 | 3.6 | 1.4 | 0.2 | 0 |
iris.target_namesarray(['setosa', 'versicolor', 'virginica'], dtype='<U10')df2.target_names.value_counts()2 50
1 50
0 50
Name: target_names, dtype: int64df2['target_names'] = df2.target_names.map({0:'setosa', 1:'versicolor', 2:'virginica'})
df2.head()| sepal length (cm) | sepal width (cm) | petal length (cm) | petal width (cm) | target_names | |
|---|---|---|---|---|---|
| 0 | 5.1 | 3.5 | 1.4 | 0.2 | setosa |
| 1 | 4.9 | 3.0 | 1.4 | 0.2 | setosa |
| 2 | 4.7 | 3.2 | 1.3 | 0.2 | setosa |
| 3 | 4.6 | 3.1 | 1.5 | 0.2 | setosa |
| 4 | 5.0 | 3.6 | 1.4 | 0.2 | setosa |
Visual EDA
_ = pd.plotting.scatter_matrix(df, c=y, figsize=[8,8], s=150, marker="D")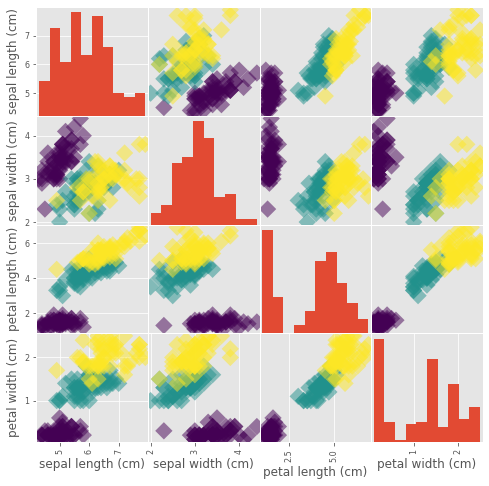
Numerical EDA
We’ll be working with a dataset obtained from the UCI Machine Learning Repository consisting of votes made by US House of Representatives Congressmen. our goal will be to predict their party affiliation (‘Democrat’ or ‘Republican’) based on how they voted on certain key issues.
Here, it’s worth noting that we have preprocessed this dataset to deal with missing values. This is so that our focus can be directed towards understanding how to train and evaluate supervised learning models.
Before thinking about what supervised learning models we can apply to this, however, we need to perform Exploratory data analysis (EDA) in order to understand the structure of the data.
votes = pd.read_csv("datasets/votes.csv")
votes.head()| party | infants | water | budget | physician | salvador | religious | satellite | aid | missile | immigration | synfuels | education | superfund | crime | duty_free_exports | eaa_rsa | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 0 | republican | 0 | 1 | 0 | 1 | 1 | 1 | 0 | 0 | 0 | 1 | 0 | 1 | 1 | 1 | 0 | 1 |
| 1 | republican | 0 | 1 | 0 | 1 | 1 | 1 | 0 | 0 | 0 | 0 | 0 | 1 | 1 | 1 | 0 | 1 |
| 2 | democrat | 0 | 1 | 1 | 0 | 1 | 1 | 0 | 0 | 0 | 0 | 1 | 0 | 1 | 1 | 0 | 0 |
| 3 | democrat | 0 | 1 | 1 | 0 | 1 | 1 | 0 | 0 | 0 | 0 | 1 | 0 | 1 | 0 | 0 | 1 |
| 4 | democrat | 1 | 1 | 1 | 0 | 1 | 1 | 0 | 0 | 0 | 0 | 1 | 0 | 1 | 1 | 1 | 1 |
votes.info()<class 'pandas.core.frame.DataFrame'>
RangeIndex: 435 entries, 0 to 434
Data columns (total 17 columns):
# Column Non-Null Count Dtype
--- ------ -------------- -----
0 party 435 non-null object
1 infants 435 non-null int64
2 water 435 non-null int64
3 budget 435 non-null int64
4 physician 435 non-null int64
5 salvador 435 non-null int64
6 religious 435 non-null int64
7 satellite 435 non-null int64
8 aid 435 non-null int64
9 missile 435 non-null int64
10 immigration 435 non-null int64
11 synfuels 435 non-null int64
12 education 435 non-null int64
13 superfund 435 non-null int64
14 crime 435 non-null int64
15 duty_free_exports 435 non-null int64
16 eaa_rsa 435 non-null int64
dtypes: int64(16), object(1)
memory usage: 57.9+ KBvotes.describe()| infants | water | budget | physician | salvador | religious | satellite | aid | missile | immigration | synfuels | education | superfund | crime | duty_free_exports | eaa_rsa | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| count | 435.000000 | 435.000000 | 435.000000 | 435.000000 | 435.000000 | 435.000000 | 435.000000 | 435.000000 | 435.000000 | 435.000000 | 435.000000 | 435.000000 | 435.000000 | 435.000000 | 435.000000 | 435.000000 |
| mean | 0.429885 | 0.558621 | 0.606897 | 0.406897 | 0.521839 | 0.650575 | 0.581609 | 0.590805 | 0.526437 | 0.512644 | 0.344828 | 0.393103 | 0.537931 | 0.609195 | 0.400000 | 0.857471 |
| std | 0.495630 | 0.497123 | 0.489002 | 0.491821 | 0.500098 | 0.477337 | 0.493863 | 0.492252 | 0.499876 | 0.500416 | 0.475859 | 0.489002 | 0.499133 | 0.488493 | 0.490462 | 0.349994 |
| min | 0.000000 | 0.000000 | 0.000000 | 0.000000 | 0.000000 | 0.000000 | 0.000000 | 0.000000 | 0.000000 | 0.000000 | 0.000000 | 0.000000 | 0.000000 | 0.000000 | 0.000000 | 0.000000 |
| 25% | 0.000000 | 0.000000 | 0.000000 | 0.000000 | 0.000000 | 0.000000 | 0.000000 | 0.000000 | 0.000000 | 0.000000 | 0.000000 | 0.000000 | 0.000000 | 0.000000 | 0.000000 | 1.000000 |
| 50% | 0.000000 | 1.000000 | 1.000000 | 0.000000 | 1.000000 | 1.000000 | 1.000000 | 1.000000 | 1.000000 | 1.000000 | 0.000000 | 0.000000 | 1.000000 | 1.000000 | 0.000000 | 1.000000 |
| 75% | 1.000000 | 1.000000 | 1.000000 | 1.000000 | 1.000000 | 1.000000 | 1.000000 | 1.000000 | 1.000000 | 1.000000 | 1.000000 | 1.000000 | 1.000000 | 1.000000 | 1.000000 | 1.000000 |
| max | 1.000000 | 1.000000 | 1.000000 | 1.000000 | 1.000000 | 1.000000 | 1.000000 | 1.000000 | 1.000000 | 1.000000 | 1.000000 | 1.000000 | 1.000000 | 1.000000 | 1.000000 | 1.000000 |
Observations
- The DataFrame has a total of 435 rows and 17 columns.
- Except for
'party', all of the columns are of typeint64. - The first two rows of the DataFrame consist of votes made by Republicans and the next three rows consist of votes made by Democrats.
- The target variable in this DataFrame is
'party'.
Votes Visual EDA
The Numerical EDA we did gave us some very important information, such as the names and data types of the columns, and the dimensions of the DataFrame. Following this with some visual EDA will give us an even better understanding of the data. all the features in this dataset are binary; that is, they are either 0 or 1. So a different type of plot would be more useful here, such as Seaborn’s countplot.
def plot_countplot(column):
plt.figure()
sns.countplot(x=column, hue='party', data=votes, palette='RdBu')
plt.xticks([0,1], ['No', 'Yes'])
plt.show()
plot_countplot("education")
It seems like Democrats voted resoundingly against this bill, compared to Republicans. This is the kind of information that our machine learning model will seek to learn when we try to predict party affiliation solely based on voting behavior. An expert in U.S politics may be able to predict this without machine learning, but probably not instantaneously - and certainly not if we are dealing with hundreds of samples!
plot_countplot('infants')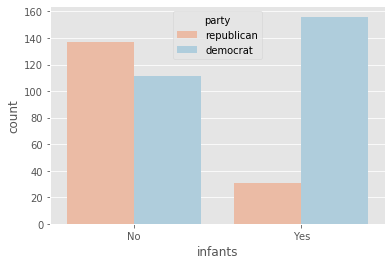
plot_countplot('water')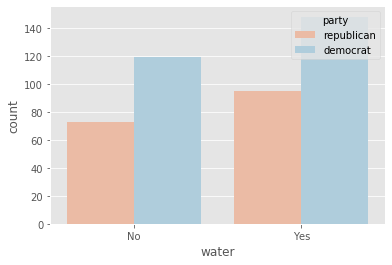
plot_countplot("budget")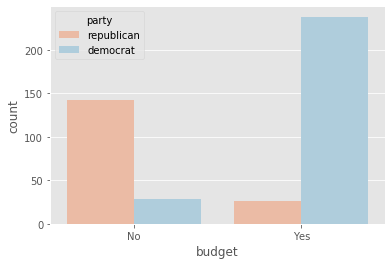
plot_countplot('physician')
plot_countplot('salvador')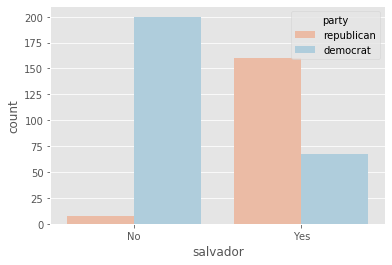
plot_countplot('religious')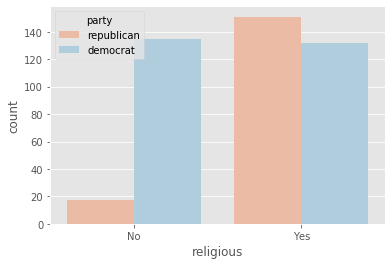
plot_countplot('satellite')
plot_countplot('aid')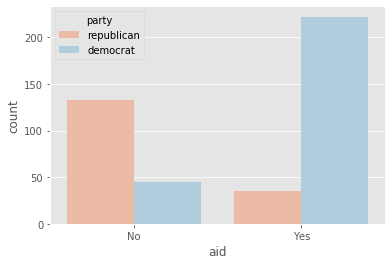
plot_countplot('missile')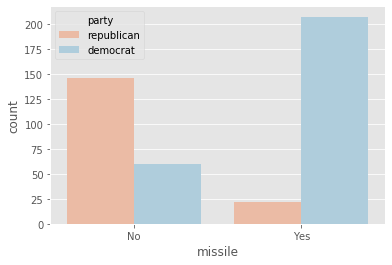
plot_countplot('immigration')
plot_countplot('synfuels')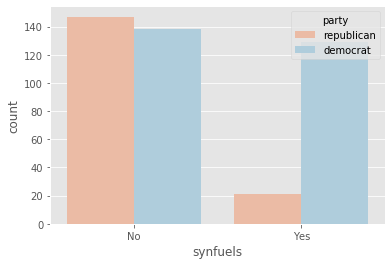
plot_countplot('superfund')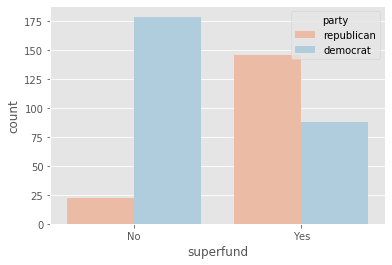
plot_countplot('crime')
plot_countplot('duty_free_exports')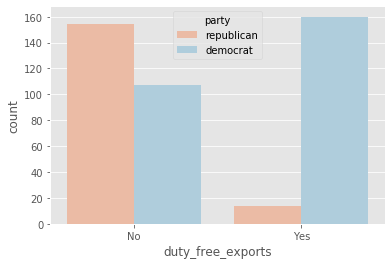
plot_countplot('eaa_rsa')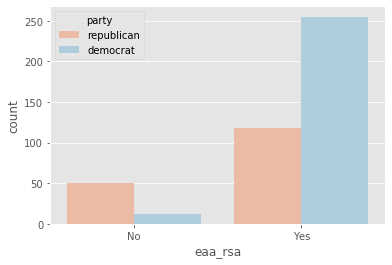
Observations
- Democrats voted in favor of both
'satellite'and'missile'
The classification challenge
k-Nearest Neighbors
- Basic idea: Predict the label of a data point by
- Looking at the ‘k’ closest labeled data points
- Taking a majority vote
Scikit-learn fit and predict
- All machine learning models implemented as Python classes
- They implement the algorithms for learning and predicting
- Store the information learned from the data
- Training a model on the data = ‘fitting’ a model to the data
.fit()method- To predict the labels of new data:
.predict()method
Iris k-NN: Intuition
_ = sns.scatterplot(data=df2, x="petal width (cm)", y="petal length (cm)", hue='target_names')
plt.show()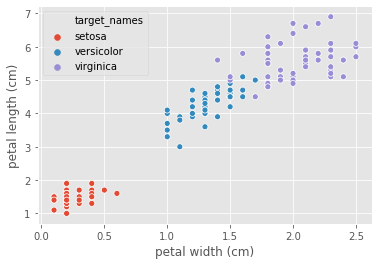
Iris dataset Using scikit-learn to fit a classier
knn = KNeighborsClassifier(n_neighbors=6)
knn.fit(iris['data'], iris['target'])KNeighborsClassifier(n_neighbors=6)iris['data'].shape(150, 4)iris['target'].shape(150,)Predicting on unlabeled data
X_new = np.array([[5.6, 2.8, 3.9, 1.1],
[5.7, 2.6, 3.8, 1.3],
[4.7, 3.2, 1.3, 0.2]])
prediction = knn.predict(X_new)
predictionarray([1, 1, 0])k-Nearest Neighbors: Fit
Having explored the Congressional voting records dataset, it is time now to build our first classifier. We’ll will fit a k-Nearest Neighbors classifier to the voting dataset.
The features need to be in an array where each column is a feature and each row a different observation or data point - in this case, a Congressman’s voting record. The target needs to be a single column with the same number of observations as the feature data. We will name the feature array X and response variable y: This is in accordance with the common scikit-learn practice.
# Create arrays for the features and the response variable
y_votes = votes['party'].values
X_votes = votes.drop('party', axis=1).values
# Create a k-NN classifier with 6 neighbors
knn_votes = KNeighborsClassifier(n_neighbors=6)
# Fit the classifier to the data
knn_votes.fit(X_votes, y_votes)KNeighborsClassifier(n_neighbors=6)Now that the k-NN classifier with 6 neighbors has been fit to the data, it can be used to predict the labels of new data points.
k-Nearest Neighbors: Predict
X_new_votes = pd.read_csv("datasets/X_new_votes.csv")
X_new_votes.head()| 0 | 1 | 2 | 3 | 4 | 5 | 6 | 7 | 8 | 9 | 10 | 11 | 12 | 13 | 14 | 15 | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 0 | 0.696469 | 0.286139 | 0.226851 | 0.551315 | 0.719469 | 0.423106 | 0.980764 | 0.68483 | 0.480932 | 0.392118 | 0.343178 | 0.72905 | 0.438572 | 0.059678 | 0.398044 | 0.737995 |
Having fit a k-NN classifier, we can now use it to predict the label of a new data point.
# Predict and print the label for the new data point X_new
new_prediction = knn_votes.predict(X_new_votes)
print("Prediction: {}".format(new_prediction))Prediction: ['democrat']Measuring model performance
- In classication, accuracy is a commonly used metric
- Accuracy = Fraction of correct predictions
- Which data should be used to compute accuracy?
- How well will the model perform on new data?
- Could compute accuracy on data used to fit classifier
- NOT indicative of ability to generalize
- Split data into training and test set
- Fit/train the classifier on the training set
- Make predictions on test set
- Compare predictions with the known labels
Model complexity
- Larger k = smoother decision boundary = less complex model
- Smaller k = more complex model = can lead to overfitting
X_train_iris, X_test_iris, y_train_iris, y_test_iris = train_test_split(X, y, test_size=.3, random_state=21, stratify=y)
knn_iris = KNeighborsClassifier(n_neighbors=8)
knn_iris.fit(X_train_iris, y_train_iris)
y_pred_iris = knn_iris.predict(X_test_iris)
print(f"Test set predictions \n{y_pred_iris}")Test set predictions
[2 1 2 2 1 0 1 0 0 1 0 2 0 2 2 0 0 0 1 0 2 2 2 0 1 1 1 0 0 1 2 2 0 0 1 2 2
1 1 2 1 1 0 2 1]knn_iris.score(X_test_iris, y_test_iris)0.9555555555555556The digits recognition dataset
We’ll be working with the MNIST digits recognition dataset, which has 10 classes, the digits 0 through 9! A reduced version of the MNIST dataset is one of scikit-learn’s included datasets.
Each sample in this scikit-learn dataset is an 8x8 image representing a handwritten digit. Each pixel is represented by an integer in the range 0 to 16, indicating varying levels of black. Helpfully for the MNIST dataset, scikit-learn provides an 'images' key in addition to the 'data' and 'target' keys that we have seen with the Iris data. Because it is a 2D array of the images corresponding to each sample, this 'images' key is useful for visualizing the images. On the other hand, the 'data' key contains the feature array - that is, the images as a flattened array of 64 pixels.
# Load the digits dataset: digits
digits = datasets.load_digits()
# Print the keys and DESCR of the dataset
print(digits.keys())
print(digits.DESCR)dict_keys(['data', 'target', 'frame', 'feature_names', 'target_names', 'images', 'DESCR'])
.. _digits_dataset:
Optical recognition of handwritten digits dataset
--------------------------------------------------
**Data Set Characteristics:**
:Number of Instances: 5620
:Number of Attributes: 64
:Attribute Information: 8x8 image of integer pixels in the range 0..16.
:Missing Attribute Values: None
:Creator: E. Alpaydin (alpaydin '@' boun.edu.tr)
:Date: July; 1998
This is a copy of the test set of the UCI ML hand-written digits datasets
https://archive.ics.uci.edu/ml/datasets/Optical+Recognition+of+Handwritten+Digits
The data set contains images of hand-written digits: 10 classes where
each class refers to a digit.
Preprocessing programs made available by NIST were used to extract
normalized bitmaps of handwritten digits from a preprinted form. From a
total of 43 people, 30 contributed to the training set and different 13
to the test set. 32x32 bitmaps are divided into nonoverlapping blocks of
4x4 and the number of on pixels are counted in each block. This generates
an input matrix of 8x8 where each element is an integer in the range
0..16. This reduces dimensionality and gives invariance to small
distortions.
For info on NIST preprocessing routines, see M. D. Garris, J. L. Blue, G.
T. Candela, D. L. Dimmick, J. Geist, P. J. Grother, S. A. Janet, and C.
L. Wilson, NIST Form-Based Handprint Recognition System, NISTIR 5469,
1994.
.. topic:: References
- C. Kaynak (1995) Methods of Combining Multiple Classifiers and Their
Applications to Handwritten Digit Recognition, MSc Thesis, Institute of
Graduate Studies in Science and Engineering, Bogazici University.
- E. Alpaydin, C. Kaynak (1998) Cascading Classifiers, Kybernetika.
- Ken Tang and Ponnuthurai N. Suganthan and Xi Yao and A. Kai Qin.
Linear dimensionalityreduction using relevance weighted LDA. School of
Electrical and Electronic Engineering Nanyang Technological University.
2005.
- Claudio Gentile. A New Approximate Maximal Margin Classification
Algorithm. NIPS. 2000.# Print the shape of the images and data keys
print(digits.images.shape)
digits.data.shape(1797, 8, 8)(1797, 64)# Display digit 1010
plt.imshow(digits.images[1010], cmap=plt.cm.gray_r, interpolation='nearest')
plt.show()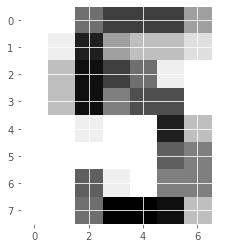
It looks like the image in question corresponds to the digit ‘5’. Now, can we build a classifier that can make this prediction not only for this image, but for all the other ones in the dataset?
Train/Test Split + Fit/Predict/Accuracy
Now that we have learned about the importance of splitting your data into training and test sets, it’s time to practice doing this on the digits dataset! After creating arrays for the features and target variable, we will split them into training and test sets, fit a k-NN classifier to the training data, and then compute its accuracy using the .score() method.
# Create feature and target arrays
X_digits = digits.data
y_digits = digits.target
# Split into training and test set
X_train_digits, X_test_digits, y_train_digits, y_test_digits = train_test_split(X_digits, y_digits,
test_size = 0.2, random_state= 42,
stratify=y_digits)
# Create a k-NN classifier with 7 neighbors: knn_digits
knn_digits = KNeighborsClassifier(n_neighbors=7)
# Fit the classifier to the training data
knn_digits.fit(X_train_digits, y_train_digits)
# Print the accuracy
knn_digits.score(X_test_digits, y_test_digits)0.9833333333333333Incredibly, this out of the box k-NN classifier with 7 neighbors has learned from the training data and predicted the labels of the images in the test set with 98% accuracy, and it did so in less than a second! This is one illustration of how incredibly useful machine learning techniques can be.
Overfitting and underfitting
We will now construct such a model complexity curve for the digits dataset! We will compute and plot the training and testing accuracy scores for a variety of different neighbor values.
By observing how the accuracy scores differ for the training and testing sets with different values of k, we will develop your intuition for overfitting and underfitting.
# Setup arrays to store train and test accuracies
neighbors_digits = np.arange(1, 9)
train_accuracy_digits = np.empty(len(neighbors_digits))
test_accuracy_digits = np.empty(len(neighbors_digits))
# Loop over different values of k
for i, k in enumerate(neighbors_digits):
# Setup a k-NN Classifier with k neighbors: knn
knn_digits = KNeighborsClassifier(n_neighbors=k)
# Fit the classifier to the training data
knn_digits.fit(X_train_digits, y_train_digits)
#Compute accuracy on the training set
train_accuracy_digits[i] = knn_digits.score(X_train_digits, y_train_digits)
#Compute accuracy on the testing set
test_accuracy_digits[i] = knn_digits.score(X_test_digits, y_test_digits)
# Generate plot
plt.title('k-NN: Varying Number of Neighbors')
plt.plot(neighbors_digits, test_accuracy_digits, label = 'Testing Accuracy')
plt.plot(neighbors_digits, train_accuracy_digits, label = 'Training Accuracy')
plt.legend()
plt.xlabel('Number of Neighbors')
plt.ylabel('Accuracy')
plt.show()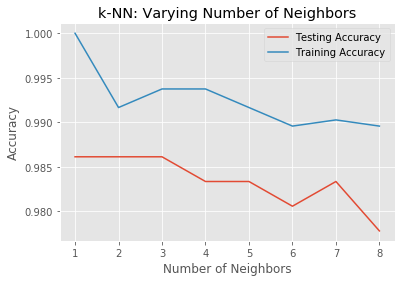
It looks like the test accuracy is highest when using 1 and35 neighbors. Using 8 neighbors or more seems to result in a simple model that underfits the data.
Regression
We used image and political datasets to predict binary and multiclass outcomes. But what if our problem requires a continuous outcome? Regression is best suited to solving such problems. We will explore the fundamental concepts in regression and apply them to predict the life expectancy in a given country using Gapminder data.
Introduction to regression
Example of an regression problem: A bike share company using time and weather data to predict the number of bikes being rented at any given hour. The target variable here - the number of bike rentals at any given hour - is quantitative, so this is best framed as a regression problem.
Boston housing data
boston = datasets.load_boston()
boston.data.shape(506, 13)boston.target.shape(506,)boston.feature_namesarray(['CRIM', 'ZN', 'INDUS', 'CHAS', 'NOX', 'RM', 'AGE', 'DIS', 'RAD',
'TAX', 'PTRATIO', 'B', 'LSTAT'], dtype='<U7')boston_df = pd.DataFrame(boston.data, columns=boston.feature_names)
boston_df['MEDV'] = boston.target
boston_df.head()| CRIM | ZN | INDUS | CHAS | NOX | RM | AGE | DIS | RAD | TAX | PTRATIO | B | LSTAT | MEDV | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 0 | 0.00632 | 18.0 | 2.31 | 0.0 | 0.538 | 6.575 | 65.2 | 4.0900 | 1.0 | 296.0 | 15.3 | 396.90 | 4.98 | 24.0 |
| 1 | 0.02731 | 0.0 | 7.07 | 0.0 | 0.469 | 6.421 | 78.9 | 4.9671 | 2.0 | 242.0 | 17.8 | 396.90 | 9.14 | 21.6 |
| 2 | 0.02729 | 0.0 | 7.07 | 0.0 | 0.469 | 7.185 | 61.1 | 4.9671 | 2.0 | 242.0 | 17.8 | 392.83 | 4.03 | 34.7 |
| 3 | 0.03237 | 0.0 | 2.18 | 0.0 | 0.458 | 6.998 | 45.8 | 6.0622 | 3.0 | 222.0 | 18.7 | 394.63 | 2.94 | 33.4 |
| 4 | 0.06905 | 0.0 | 2.18 | 0.0 | 0.458 | 7.147 | 54.2 | 6.0622 | 3.0 | 222.0 | 18.7 | 396.90 | 5.33 | 36.2 |
Creating feature and target arrays for the boston dataset
X_boston = boston.data
y_boston = boston.targetPredicting house value from a single feature
X_boston_rooms = X_boston[:,5]
type(X_boston_rooms), type(y_boston)(numpy.ndarray, numpy.ndarray)y_boston = y_boston.reshape(-1,1)
X_boston_rooms = X_boston_rooms.reshape(-1,1)Plotting house value vs. number of rooms
plt.scatter(X_boston_rooms, y_boston)
plt.ylabel('Value of house /1000 ($)')
plt.xlabel('Number of rooms')
plt.show();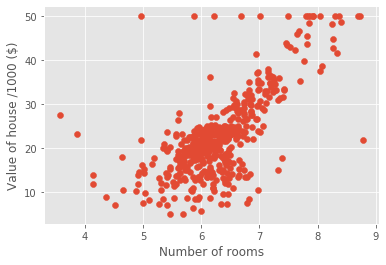
Fitting a regression model
reg_boston = LinearRegression()
reg_boston.fit(X_boston_rooms, y_boston)
boston_prediction_space = np.linspace(min(X_boston_rooms), max(X_boston_rooms)).reshape(-1,1)plt.scatter(X_boston_rooms, y_boston, color="blue")
plt.plot(boston_prediction_space, reg_boston.predict(boston_prediction_space), color='black', linewidth=3)
plt.show()
Importing Gapminder data for supervised learning
We will work with Gapminder data that we have consolidated into one CSV file.
Specifically, our goal will be to use this data to predict the life expectancy in a given country based on features such as the country’s GDP, fertility rate, and population.
Since the target variable here is quantitative, this is a regression problem. To begin, we will fit a linear regression with just one feature: 'fertility', which is the average number of children a woman in a given country gives birth to.
Before that, however, we need to import the data and get it into the form needed by scikit-learn. This involves creating feature and target variable arrays. Furthermore, since we are going to use only one feature to begin with, we need to do some reshaping using NumPy’s .reshape() method.
# Read the CSV file into a DataFrame: gapminder_df
gapminder = pd.read_csv("datasets/gapminder.csv")
# Create arrays for features and target variable
y_gapminder = gapminder.life.values
X_gapminder = gapminder.fertility.values
# Print the dimensions of X and y before reshaping
print("Dimensions of y before reshaping: {}".format(y_gapminder.shape))
print("Dimensions of X before reshaping: {}".format(X_gapminder.shape))
# Reshape X and y
y_gapminder = y_gapminder.reshape(-1,1)
X_gapminder = X_gapminder.reshape(-1,1)
# Print the dimensions of X and y after reshaping
print("Dimensions of y after reshaping: {}".format(y_gapminder.shape))
print("Dimensions of X after reshaping: {}".format(X_gapminder.shape))Dimensions of y before reshaping: (139,)
Dimensions of X before reshaping: (139,)
Dimensions of y after reshaping: (139, 1)
Dimensions of X after reshaping: (139, 1)Exploring the Gapminder data
As always, it is important to explore the data before building models.
sns.heatmap(gapminder.corr(), square=True, cmap="RdYlGn")
plt.show()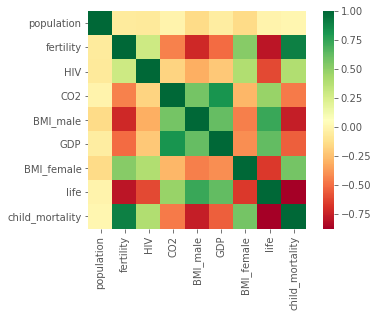
Cells that are in green show positive correlation, while cells that are in red show negative correlation. life and fertility are negatively correlated. GDP and life are positively correlated
gapminder.head()| population | fertility | HIV | CO2 | BMI_male | GDP | BMI_female | life | child_mortality | |
|---|---|---|---|---|---|---|---|---|---|
| 0 | 34811059.0 | 2.73 | 0.1 | 3.328945 | 24.59620 | 12314.0 | 129.9049 | 75.3 | 29.5 |
| 1 | 19842251.0 | 6.43 | 2.0 | 1.474353 | 22.25083 | 7103.0 | 130.1247 | 58.3 | 192.0 |
| 2 | 40381860.0 | 2.24 | 0.5 | 4.785170 | 27.50170 | 14646.0 | 118.8915 | 75.5 | 15.4 |
| 3 | 2975029.0 | 1.40 | 0.1 | 1.804106 | 25.35542 | 7383.0 | 132.8108 | 72.5 | 20.0 |
| 4 | 21370348.0 | 1.96 | 0.1 | 18.016313 | 27.56373 | 41312.0 | 117.3755 | 81.5 | 5.2 |
gapminder.info()<class 'pandas.core.frame.DataFrame'>
RangeIndex: 139 entries, 0 to 138
Data columns (total 9 columns):
# Column Non-Null Count Dtype
--- ------ -------------- -----
0 population 139 non-null float64
1 fertility 139 non-null float64
2 HIV 139 non-null float64
3 CO2 139 non-null float64
4 BMI_male 139 non-null float64
5 GDP 139 non-null float64
6 BMI_female 139 non-null float64
7 life 139 non-null float64
8 child_mortality 139 non-null float64
dtypes: float64(9)
memory usage: 9.9 KBThe DataFrame has 139 samples (or rows) and 9 columns.
gapminder.describe()| population | fertility | HIV | CO2 | BMI_male | GDP | BMI_female | life | child_mortality | |
|---|---|---|---|---|---|---|---|---|---|
| count | 1.390000e+02 | 139.000000 | 139.000000 | 139.000000 | 139.000000 | 139.000000 | 139.000000 | 139.000000 | 139.000000 |
| mean | 3.549977e+07 | 3.005108 | 1.915612 | 4.459874 | 24.623054 | 16638.784173 | 126.701914 | 69.602878 | 45.097122 |
| std | 1.095121e+08 | 1.615354 | 4.408974 | 6.268349 | 2.209368 | 19207.299083 | 4.471997 | 9.122189 | 45.724667 |
| min | 2.773150e+05 | 1.280000 | 0.060000 | 0.008618 | 20.397420 | 588.000000 | 117.375500 | 45.200000 | 2.700000 |
| 25% | 3.752776e+06 | 1.810000 | 0.100000 | 0.496190 | 22.448135 | 2899.000000 | 123.232200 | 62.200000 | 8.100000 |
| 50% | 9.705130e+06 | 2.410000 | 0.400000 | 2.223796 | 25.156990 | 9938.000000 | 126.519600 | 72.000000 | 24.000000 |
| 75% | 2.791973e+07 | 4.095000 | 1.300000 | 6.589156 | 26.497575 | 23278.500000 | 130.275900 | 76.850000 | 74.200000 |
| max | 1.197070e+09 | 7.590000 | 25.900000 | 48.702062 | 28.456980 | 126076.000000 | 135.492000 | 82.600000 | 192.000000 |
The mean of life is 69.602878
The basics of linear regression
Regression mechanics
- \(y = ax + b\)
- \(y\) = target
- \(x\) = single feature
- \(a\), \(b\) = parameters of model
- How do we choose \(a\) and \(b\)?
- Define an error functions for any given lineChoose the line that minimizes the error function
- Ordinary least squares(OLS): Minimize sum of squares of residuals
Linear regression in higher dimensions
- \(y=a_1x_1+a_2x_2+b\)
- To fit a linear regression model here:
- Need to specify 3 variables
- In higher dimensions:
- Must specify coefcient for each feature and the variable \(b\)
- \(y=a_1x_1+a_2x_2+a_3x_3+...+a_nx_n+b\)
- Scikit-learn API works exactly the same way:
- Pass two arrays: Features, and target
Linear regression on all features in boston dataset
X_train_boston, X_test_boston, y_train_boston, y_test_boston = train_test_split(X_boston, y_boston,
test_size=.3, random_state=42)
reg_all_boston = LinearRegression()
reg_all_boston.fit(X_train_boston, y_train_boston)
y_pred_boston = reg_all_boston.predict(X_test_boston)
reg_all_boston.score(X_test_boston, y_test_boston)0.7112260057484925Fit & predict for regression in gapminder dataset
We will fit a linear regression and predict life expectancy using just one feature. We will use the 'fertility' feature of the Gapminder dataset. Since the goal is to predict life expectancy, the target variable here is 'life'.
sns.scatterplot(data=gapminder, x="fertility", y="life")
plt.show()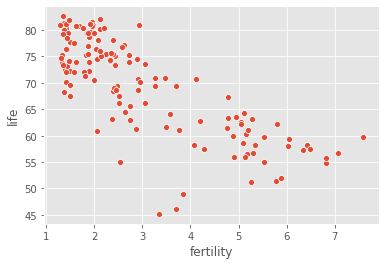
As you can see, there is a strongly negative correlation, so a linear regression should be able to capture this trend. Our job is to fit a linear regression and then predict the life expectancy, overlaying these predicted values on the plot to generate a regression line. We will also compute and print the \(R^2\) score using sckit-learn’s .score() method.
# Create the regressor: reg
reg_gapminder = LinearRegression()
# Create the prediction space
prediction_space = np.linspace(min(X_gapminder), max(X_gapminder)).reshape(-1,1)
# Fit the model to the data
reg_gapminder.fit(X_gapminder,y_gapminder)
# Compute predictions over the prediction space: y_pred
y_pred_gapminder = reg_gapminder.predict(prediction_space)
# Print R^2
print(reg_gapminder.score(X_gapminder, y_gapminder))0.6192442167740035# Plot regression line
sns.scatterplot(data=gapminder, x="fertility", y="life")
plt.plot(prediction_space, y_pred_gapminder, color='black', linewidth=3)
plt.show()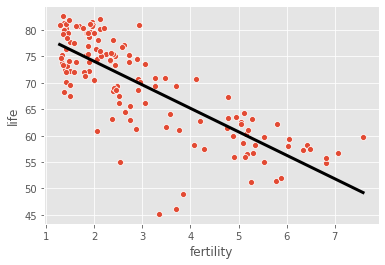
Notice how the line captures the underlying trend in the data. And the performance is quite decent for this basic regression model with only one feature!
Train/test split for regression
train and test sets are vital to ensure that the supervised learning model is able to generalize well to new data. This was true for classification models, and is equally true for linear regression models.
We will split the Gapminder dataset into training and testing sets, and then fit and predict a linear regression over all features. In addition to computing the \(R^2\) score, we will also compute the Root Mean Squared Error (RMSE), which is another commonly used metric to evaluate regression models.
X_gapminder = gapminder.drop("life", axis=1).values# Create training and test sets
X_train_gapminder, X_test_gapminder, y_train_gapminder, y_test_gapminder = train_test_split(X_gapminder, y_gapminder, test_size = .3, random_state=42)
# Create the regressor: reg_all
reg_all_gapminder = LinearRegression()
# Fit the regressor to the training data
reg_all_gapminder.fit(X_train_gapminder, y_train_gapminder)
# Predict on the test data: y_pred
y_pred_gapminder = reg_all_gapminder.predict(X_test_gapminder)
# Compute and print R^2 and RMSE
print("R^2: {}".format(reg_all_gapminder.score(X_test_gapminder, y_test_gapminder)))
rmse_gapminder = np.sqrt(mean_squared_error(y_test_gapminder, y_pred_gapminder))
print("Root Mean Squared Error: {}".format(rmse_gapminder))R^2: 0.8380468731430059
Root Mean Squared Error: 3.247601080037022Using all features has improved the model score. This makes sense, as the model has more information to learn from. However, there is one potential pitfall to this process. Can you spot it?
Cross-validation
Cross-validation motivation
- Model performance is dependent on way the data is split
- Not representative of the model’s ability to generalize
- Solution: Cross-validation!
Cross-validation and model performance
- 5 folds = 5-fold CV
- 10 folds = 10-fold CV
- k folds = k-fold CV
- More folds = More computationally expensive
Cross-validation in scikit-learn: Boston
cv_results_boston = cross_val_score(reg_all_boston, X_boston, y_boston, cv=5)
cv_results_bostonarray([ 0.63919994, 0.71386698, 0.58702344, 0.07923081, -0.25294154])np.mean(cv_results_boston)0.353275924395884np.median(cv_results_boston)0.58702343630577765-fold cross-validation
Cross-validation is a vital step in evaluating a model. It maximizes the amount of data that is used to train the model, as during the course of training, the model is not only trained, but also tested on all of the available data.
We will practice 5-fold cross validation on the Gapminder data. By default, scikit-learn’s cross_val_score() function uses R2 as the metric of choice for regression. Since We are performing 5-fold cross-validation, the function will return 5 scores. We will compute these 5 scores and then take their average.
# Compute 5-fold cross-validation scores: cv_scores
cv_scores_gapminder = cross_val_score(reg_gapminder, X_gapminder, y_gapminder, cv=5)
# Print the 5-fold cross-validation scores
print(cv_scores_gapminder)
print("Average 5-Fold CV Score: {}".format(np.mean(cv_scores_gapminder)))[0.81720569 0.82917058 0.90214134 0.80633989 0.94495637]
Average 5-Fold CV Score: 0.8599627722793267Now that we have cross-validated your model, we can more confidently evaluate its predictions.
K-Fold CV comparison
Cross validation is essential but do not forget that the more folds you use, the more computationally expensive cross-validation becomes.
%timeit cross_val_score(reg_gapminder, X_gapminder, y_gapminder, cv=3)8.03 ms ± 1.01 ms per loop (mean ± std. dev. of 7 runs, 100 loops each)%timeit cross_val_score(reg_gapminder, X_gapminder, y_gapminder, cv=10)31.8 ms ± 1.21 ms per loop (mean ± std. dev. of 7 runs, 10 loops each)# Perform 3-fold CV
cvscores_3_gapminder = cross_val_score(reg_gapminder, X_gapminder, y_gapminder, cv=3)
print(np.mean(cvscores_3_gapminder))
# Perform 10-fold CV
cvscores_10_gapminder = cross_val_score(reg_gapminder, X_gapminder, y_gapminder, cv=10)
print(np.mean(cvscores_10_gapminder))0.8718712782621969
0.8436128620131095Regularized regression
Why regularize?
- Recall: Linear regression minimizes a loss function
- It chooses a coefcient for each feature variable
- Large coefcients can lead to overtting
- Penalizing large coefcients: Regularization
Ridge regression
- Loss function = OLS loss function + \(\alpha * \sum_{i=1}^{n} a_i^2\)
- Alpha: Parameter we need to choose
- Picking alpha here is similar to picking k in k-NN
- Hyperparameter tuning
- Alpha controls model complexity
- Alpha = 0: We get back OLS (Can lead to overtting)
- Very high alpha: Can lead to undertting
Lasso regression
- Loss function = OLS loss function + \(\alpha * \sum_{i=1}^{n} |a_i|\)
Lasso regression for feature selection
- Can be used to select important features of a dataset
- Shrinks the coefcients of less important features to exactly 0
Ridge regression in scikit-learn: Boston
ridge_boston = Ridge(alpha=.1, normalize=True)
ridge_boston.fit(X_train_boston, y_train_boston)
ridge_pred_boston = ridge_boston.predict(X_test_boston)
ridge_boston.score(X_test_boston, y_test_boston)0.6996938275127311Lasso regression in scikit-learn: Boston
lasso_boston = Lasso(alpha=.1, normalize=True)
lasso_boston.fit(X_train_boston, y_train_boston)
lasso_pred_boston = lasso_boston.predict(X_test_boston)
lasso_boston.score(X_test_boston, y_test_boston)0.5950229535328551Lasso for feature selection in scikit-learn: Boston
names_boston = boston.feature_names
lasso_boston_2 = Lasso(alpha=.1)
lasso_coef_boston = lasso_boston_2.fit(X_boston, y_boston).coef_
_ = plt.plot(range(len(names_boston)), lasso_coef_boston)
_ = plt.xticks(range(len(names_boston)), names_boston, rotation=60)
_ = plt.ylabel("Coefficients")
plt.show()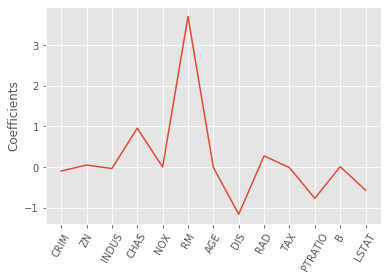
Regularization I: Lasso
We saw how Lasso selected out the ‘RM’ feature as being the most important for predicting Boston house prices, while shrinking the coefficients of certain other features to 0. Its ability to perform feature selection in this way becomes even more useful when you are dealing with data involving thousands of features.
We will fit a lasso regression to the Gapminder data we have been working with and plot the coefficients. Just as with the Boston data.
df_columns_gapminder = pd.Index(['population', 'fertility', 'HIV', 'CO2', 'BMI_male', 'GDP',
'BMI_female', 'child_mortality'],
dtype='object')# Instantiate a lasso regressor: lasso
lasso_gapminder = Lasso(alpha=.4, normalize=True)
# Fit the regressor to the data
lasso_gapminder.fit(X_gapminder,y_gapminder)
# Compute and print the coefficients
lasso_coef_gapminder = lasso_gapminder.fit(X_gapminder,y_gapminder).coef_
print(lasso_coef_gapminder)
# Plot the coefficients
plt.plot(range(len(df_columns_gapminder)), lasso_coef_gapminder)
plt.xticks(range(len(df_columns_gapminder)), df_columns_gapminder.values, rotation=60)
plt.margins(0.02)
plt.show()[-0. -0. -0. 0. 0. 0.
-0. -0.07087587]
According to the lasso algorithm, it seems like 'child_mortality' is the most important feature when predicting life expectancy.
Regularization II: Ridge
Lasso is great for feature selection, but when building regression models, Ridge regression should be the first choice.
lasso performs regularization by adding to the loss function a penalty term of the absolute value of each coefficient multiplied by some alpha. This is also known as \(L1\) regularization because the regularization term is the \(L1\) norm of the coefficients. This is not the only way to regularize, however.
def display_plot(cv_scores, cv_scores_std):
"""plots the R^2 score as well as standard error for each alpha"""
fig = plt.figure()
ax = fig.add_subplot(1,1,1)
ax.plot(alpha_space_gapminder, cv_scores)
std_error = cv_scores_std / np.sqrt(10)
ax.fill_between(alpha_space_gapminder, cv_scores + std_error, cv_scores - std_error, alpha=0.2)
ax.set_ylabel('CV Score +/- Std Error')
ax.set_xlabel('Alpha')
ax.axhline(np.max(cv_scores), linestyle='--', color='.5')
ax.set_xlim([alpha_space_gapminder[0], alpha_space_gapminder[-1]])
ax.set_xscale('log')
plt.show()If instead we took the sum of the squared values of the coefficients multiplied by some alpha - like in Ridge regression - we would be computing the \(L2\) norm. We will fit ridge regression models over a range of different alphas, and plot cross-validated \(R^2\) scores for each, using this function display_plot, which plots the \(R^2\) score as well as standard error for each alpha:
# Setup the array of alphas and lists to store scores
alpha_space_gapminder = np.logspace(-4, 0, 50)
ridge_scores_gapminder = []
ridge_scores_std_gapminder = []
# Create a ridge regressor: ridge
ridge_gapminder = Ridge(normalize=True)
# Compute scores over range of alphas
for alpha in alpha_space_gapminder:
# Specify the alpha value to use: ridge.alpha
ridge_gapminder.alpha = alpha
# Perform 10-fold CV: ridge_cv_scores
ridge_cv_scores_gapminder = cross_val_score(ridge_gapminder, X_gapminder, y_gapminder, cv=10)
# Append the mean of ridge_cv_scores to ridge_scores
ridge_scores_gapminder.append(np.mean(ridge_cv_scores_gapminder))
# Append the std of ridge_cv_scores to ridge_scores_std
ridge_scores_std_gapminder.append(np.std(ridge_cv_scores_gapminder))
# Display the plot
display_plot(ridge_scores_gapminder, ridge_scores_std_gapminder)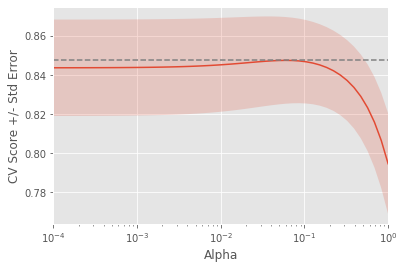
the cross-validation scores change with different alphas.
Fine-tuning model
Having trained a model, the next task is to evaluate its performance. We will explore some of the other metrics available in scikit-learn that will allow us to assess the model’s performance in a more nuanced manner.
How good is your model?
Classication metrics
- Measuring model performance with accuracy:
- Fraction of correctly classied samples
- Not always a useful metric
Class imbalance example: Emails
- Spam classication
- 99% of emails are real; 1% of emails are spam
- Could build a classier that predicts ALL emails as real
- 99% accurate!
- But horrible at actually classifying spam
- Fails at its original purpose
- Need more nuanced metrics
Diagnosing classication predictions
- Confusion matrix
- Accuracy: \(\frac{tp+tn}{tp+tn+fp+fn}\)
Metrics from the confusion matrix
- Precision: \(\frac{tp}{tp+fp}\)
- Recal \(\frac{tp}{tp+fn}\)
- F1score: \(2.\frac{precision.recal}{precision+recall}\)
- High precision: Not many real emails predicted as spam
- High recall: Predicted most spam emails correctly
Confusion matrix in scikit-learn: iris dataset
confusion_matrix(y_test_iris, y_pred_iris)array([[15, 0, 0],
[ 0, 14, 1],
[ 0, 1, 14]], dtype=int64)print(classification_report(y_test_iris, y_pred_iris)) precision recall f1-score support
0 1.00 1.00 1.00 15
1 0.93 0.93 0.93 15
2 0.93 0.93 0.93 15
accuracy 0.96 45
macro avg 0.96 0.96 0.96 45
weighted avg 0.96 0.96 0.96 45
X_train_votes, X_test_votes, y_train_votes, y_test_votes = train_test_split(X_votes, y_votes, test_size=.4, random_state=42)
knn_votes = KNeighborsClassifier(n_neighbors=8)
knn_votes.fit(X_train_votes, y_train_votes)
y_pred_votes = knn_votes.predict(X_test_votes)confusion_matrix(y_test_votes, y_pred_votes)array([[108, 7],
[ 4, 55]], dtype=int64)print(classification_report(y_test_votes, y_pred_votes)) precision recall f1-score support
democrat 0.96 0.94 0.95 115
republican 0.89 0.93 0.91 59
accuracy 0.94 174
macro avg 0.93 0.94 0.93 174
weighted avg 0.94 0.94 0.94 174
The support gives the number of samples of the true response that lie in that class, the support was the number of Republicans or Democrats in the test set on which the classification report was computed. The precision, recall, and f1-score columns, then, gave the respective metrics for that particular class.
Metrics for classification
We evaluated the performance of k-NN classifier based on its accuracy. However, accuracy is not always an informative metric. We will dive more deeply into evaluating the performance of binary classifiers by computing a confusion matrix and generating a classification report.
We’ll work with the PIMA Indians dataset obtained from the UCI Machine Learning Repository. The goal is to predict whether or not a given female patient will contract diabetes based on features such as BMI, age, and number of pregnancies. Therefore, it is a binary classification problem. A target value of 0 indicates that the patient does not have diabetes, while a value of 1 indicates that the patient does have diabetes.
pidd = pd.read_csv("datasets/pima_indians_diabetes_database.csv")
pidd.head()| pregnancies | glucose | diastolic | triceps | insulin | bmi | dpf | age | diabetes | |
|---|---|---|---|---|---|---|---|---|---|
| 0 | 6 | 148 | 72 | 35.00000 | 155.548223 | 33.6 | 0.627 | 50 | 1 |
| 1 | 1 | 85 | 66 | 29.00000 | 155.548223 | 26.6 | 0.351 | 31 | 0 |
| 2 | 8 | 183 | 64 | 29.15342 | 155.548223 | 23.3 | 0.672 | 32 | 1 |
| 3 | 1 | 89 | 66 | 23.00000 | 94.000000 | 28.1 | 0.167 | 21 | 0 |
| 4 | 0 | 137 | 40 | 35.00000 | 168.000000 | 43.1 | 2.288 | 33 | 1 |
We will train a k-NN classifier to the data and evaluate its performance by generating a confusion matrix and classification report.
y_pidd = pidd.diabetes.values
X_pidd = pidd.drop("diabetes", axis=1).values# Create training and test set
X_train_pidd, X_test_pidd, y_train_pidd, y_test_pidd = train_test_split(X_pidd, y_pidd, test_size=.4, random_state=42)
# Instantiate a k-NN classifier: knn
knn_pidd = KNeighborsClassifier(n_neighbors=6)
# Fit the classifier to the training data
knn_pidd.fit(X_train_pidd, y_train_pidd)
# Predict the labels of the test data: y_pred
y_pred_pidd = knn_pidd.predict(X_test_pidd)
# Generate the confusion matrix and classification report
print(confusion_matrix(y_test_pidd, y_pred_pidd))
print(classification_report(y_test_pidd, y_pred_pidd))[[176 30]
[ 52 50]]
precision recall f1-score support
0 0.77 0.85 0.81 206
1 0.62 0.49 0.55 102
accuracy 0.73 308
macro avg 0.70 0.67 0.68 308
weighted avg 0.72 0.73 0.72 308
By analyzing the confusion matrix and classification report, we can get a much better understanding of your classifier’s performance.
Logistic regression and the ROC curve
Logistic regression for binary classication
- Logistic regression outputs probabilities
- If the probability ‘p’ is greater than 0.5:
- The data is labeled ‘1’
- If the probability ‘p’ is less than 0.5:
- The data is labeled ‘0’
Probability thresholds
- By default, logistic regression threshold = 0.5
- Not specific to logistic regression
- k-NN classifiers also have thresholds
- What happens if we vary the threshold?
Building a logistic regression model
Time to build our first logistic regression model! scikit-learn makes it very easy to try different models, since the Train-Test-Split/Instantiate/Fit/Predict paradigm applies to all classifiers and regressors - which are known in scikit-learn as ‘estimators’.
# Create the classifier: logreg
logreg_pidd = LogisticRegression()
# Fit the classifier to the training data
logreg_pidd.fit(X_train_pidd, y_train_pidd)
# Predict the labels of the test set: y_pred
y_pred_logreg_pidd = logreg_pidd.predict(X_test_pidd)
# Compute and print the confusion matrix and classification report
print(confusion_matrix(y_test_pidd, y_pred_logreg_pidd))
print(classification_report(y_test_pidd, y_pred_logreg_pidd))[[171 35]
[ 35 67]]
precision recall f1-score support
0 0.83 0.83 0.83 206
1 0.66 0.66 0.66 102
accuracy 0.77 308
macro avg 0.74 0.74 0.74 308
weighted avg 0.77 0.77 0.77 308
Precision-recall Curve
the precision-recall curve is generated by plotting the precision and recall for different thresholds.
\[ precision = \frac{TP}{TP+FP} \] \[ recall = \frac{TP}{TP+FN} \]
disp = plot_precision_recall_curve(logreg_pidd, X_test_pidd, y_test_pidd)
disp.ax_.set_title('Precision-Recall curve: ')Text(0.5, 1.0, 'Precision-Recall curve: ')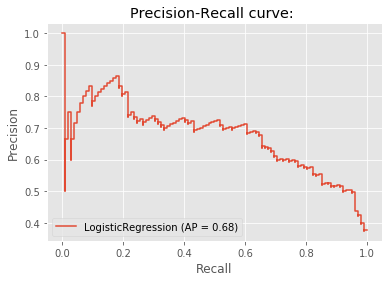
- A recall of 1 corresponds to a classifier with a low threshold in which all females who contract diabetes were correctly classified as such, at the expense of many misclassifications of those who did not have diabetes.
- Precision is undefined for a classifier which makes no positive predictions, that is, classifies everyone as not having diabetes.
- When the threshold is very close to 1, precision is also 1, because the classifier is absolutely certain about its predictions.
Plotting an ROC curve
Classification reports and confusion matrices are great methods to quantitatively evaluate model performance, while ROC curves provide a way to visually evaluate models. most classifiers in scikit-learn have a .predict_proba() method which returns the probability of a given sample being in a particular class. Having built a logistic regression model, we’ll now evaluate its performance by plotting an ROC curve. In doing so, we’ll make use of the .predict_proba() method and become familiar with its functionality.
# Compute predicted probabilities: y_pred_prob
y_pred_prob_pidd = logreg_pidd.predict_proba(X_test_pidd)[:,1]
# Generate ROC curve values: fpr, tpr, thresholds
fpr_pidd, tpr_pidd, thresholds_pidd = roc_curve(y_test_pidd, y_pred_prob_pidd)
# Plot ROC curve
plt.plot([0, 1], [0, 1], 'k--')
plt.plot(fpr_pidd, tpr_pidd)
plt.xlabel('False Positive Rate')
plt.ylabel('True Positive Rate')
plt.title('ROC Curve')
plt.show()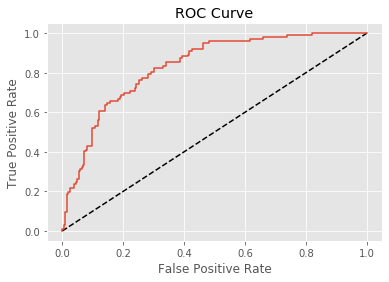
Area under the ROC curve
Area under the ROC curve (AUC)
- Larger area under the ROC curve = better model
AUC computation
Say you have a binary classifier that in fact is just randomly making guesses. It would be correct approximately 50% of the time, and the resulting ROC curve would be a diagonal line in which the True Positive Rate and False Positive Rate are always equal. The Area under this ROC curve would be 0.5. This is one way in which the AUC is an informative metric to evaluate a model. If the AUC is greater than 0.5, the model is better than random guessing. Always a good sign!
We’ll calculate AUC scores using the roc_auc_score() function from sklearn.metrics as well as by performing cross-validation on the diabetes dataset.
# Compute and print AUC score
print("AUC: {}".format(roc_auc_score(y_test_pidd, y_pred_prob_pidd)))
# Compute cross-validated AUC scores: cv_auc
cv_auc_pidd = cross_val_score(logreg_pidd, X_pidd, y_pidd, cv=5, scoring="roc_auc")
# Print list of AUC scores
print("AUC scores computed using 5-fold cross-validation: {}".format(cv_auc_pidd))AUC: 0.8329050066628594
AUC scores computed using 5-fold cross-validation: [0.81962963 0.80537037 0.82555556 0.87377358 0.82509434]Hyperparameter tuning
Hyperparameter tuning
- Linear regression: Choosing parameters
- Ridge/lasso regression: Choosing alpha
- k-Nearest Neighbors: Choosing n_neighbors
- Parameters like alpha and k: Hyperparameters
- Hyperparameters cannot be learned by tting the model
Choosing the correct hyperparameter
- Try a bunch of different hyperparameter values
- Fit all of them separately
- See how well each performs
- Choose the best performing one
- It is essential to use cross-validation
GridSearchCV in scikit-learn votes dataset
param_grid_votes = {"n_neighbors":np.arange(1,50)}
knn_votes = KNeighborsClassifier()
knn_cv_votes = GridSearchCV(knn_votes, param_grid=param_grid_votes, cv=5)
knn_cv_votes.fit(X_votes, y_votes)
knn_cv_votes.best_params_{'n_neighbors': 4}knn_cv_votes.best_score_0.9333333333333333Hyperparameter tuning with GridSearchCV
logistic regression also has a regularization parameter: \(C\). \(C\) controls the inverse of the regularization strength, and this is what we will tune. A large \(C\) can lead to an overfit model, while a small \(C\) can lead to an underfit model.
# Setup the hyperparameter grid
c_space_pidd = np.logspace(-5, 8, 15)
param_grid_pidd = {'C': c_space_pidd}
# Instantiate the GridSearchCV object: logreg_cv
logreg_cv_pidd = GridSearchCV(logreg_pidd, param_grid_pidd, cv=5)
# Fit it to the data
logreg_cv_pidd.fit(X_pidd,y_pidd)
# Print the tuned parameters and score
print("Tuned Logistic Regression Parameters: {}".format(logreg_cv_pidd.best_params_))
print("Best score is {}".format(logreg_cv_pidd.best_score_))Tuned Logistic Regression Parameters: {'C': 1389495.494373136}
Best score is 0.7787029963500551Hyperparameter tuning with RandomizedSearchCV
GridSearchCV can be computationally expensive, especially if you are searching over a large hyperparameter space and dealing with multiple hyperparameters. A solution to this is to use RandomizedSearchCV, in which not all hyperparameter values are tried out. Instead, a fixed number of hyperparameter settings is sampled from specified probability distributions.
Decision trees have many parameters that can be tuned, such as max_features, max_depth, and min_samples_leaf: This makes it an ideal use case for RandomizedSearchCV. Our goal is to use RandomizedSearchCV to find the optimal hyperparameters.
# Setup the parameters and distributions to sample from: param_dist
param_dist_pidd = {"max_depth": [3, None],
"max_features": randint(1, 9),
"min_samples_leaf": randint(1, 9),
"criterion": ["gini", "entropy"]}
# Instantiate a Decision Tree classifier: tree
tree_pidd = DecisionTreeClassifier()
# Instantiate the RandomizedSearchCV object: tree_cv
tree_cv_pidd = RandomizedSearchCV(tree_pidd, param_dist_pidd, cv=5)
# Fit it to the data
tree_cv_pidd.fit(X,y)
# Print the tuned parameters and score
print("Tuned Decision Tree Parameters: {}".format(tree_cv_pidd.best_params_))
print("Best score is {}".format(tree_cv_pidd.best_score_))Tuned Decision Tree Parameters: {'criterion': 'gini', 'max_depth': 3, 'max_features': 3, 'min_samples_leaf': 2}
Best score is 0.96RandomizedSearchCV will never outperform GridSearchCV. Instead, it is valuable because it saves on computation time.
Hold-out set for final evaluation
Hold-out set reasoning
- How well can the model perform on never before seen data?
- Using ALL data for cross-validation is not ideal
- Split data into training and hold-out set at the beginning
- Perform grid search cross-validation on training set
- Choose best hyperparameters and evaluate on hold-out set
Hold-out set in practice I: Classification
You will now practice evaluating a model with tuned hyperparameters on a hold-out set. In addition to \(C\), logistic regression has a 'penalty' hyperparameter which specifies whether to use 'l1' or 'l2' regularization. Our job is to create a hold-out set, tune the 'C' and 'penalty' hyperparameters of a logistic regression classifier using GridSearchCV on the training set.
param_grid_pidd['penalty'] = ['l1', 'l2']
# Instantiate the GridSearchCV object: logreg_cv
logreg_cv_pidd = GridSearchCV(logreg_pidd, param_grid_pidd, cv=5)
# Fit it to the training data
logreg_cv_pidd.fit(X_train_pidd, y_train_pidd)
# Print the optimal parameters and best score
print("Tuned Logistic Regression Parameter: {}".format(logreg_cv_pidd.best_params_))
print("Tuned Logistic Regression Accuracy: {}".format(logreg_cv_pidd.best_score_))Tuned Logistic Regression Parameter: {'C': 100000000.0, 'penalty': 'l2'}
Tuned Logistic Regression Accuracy: 0.7717391304347827Hold-out set in practice II: Regression
Lasso used the \(L1\) penalty to regularize, while ridge used the \(L2\) penalty. There is another type of regularized regression known as the elastic net. In elastic net regularization, the penalty term is a linear combination of the \(L1\) and \(L2\) penalties:
\[ a∗L1+b∗L2 \]
In scikit-learn, this term is represented by the 'l1_ratio' parameter: An 'l1_ratio' of 1 corresponds to an \(L1\) penalty, and anything lower is a combination of \(L1\) and \(L2\).
We will GridSearchCV to tune the 'l1_ratio' of an elastic net model trained on the Gapminder data.
# Create the hyperparameter grid
l1_space_gapminder = np.linspace(0, 1, 30)
param_grid_gapminder = {'l1_ratio': l1_space_gapminder}
# Instantiate the ElasticNet regressor: elastic_net
elastic_net_gapminder = ElasticNet()
# Setup the GridSearchCV object: gm_cv
gm_cv_gapminder = GridSearchCV(elastic_net_gapminder, param_grid_gapminder, cv=5)
# Fit it to the training data
gm_cv_gapminder.fit(X_train_gapminder, y_train_gapminder)
# Predict on the test set and compute metrics
y_pred_gapminder = gm_cv_gapminder.predict(X_test_gapminder)
r2_gapminder = gm_cv_gapminder.score(X_test_gapminder, y_test_gapminder)
mse_gapminder = mean_squared_error(y_test_gapminder, y_pred_gapminder)
print("Tuned ElasticNet l1 ratio: {}".format(gm_cv_gapminder.best_params_))
print("Tuned ElasticNet R squared: {}".format(r2_gapminder))
print("Tuned ElasticNet MSE: {}".format(mse_gapminder))Tuned ElasticNet l1 ratio: {'l1_ratio': 0.0}
Tuned ElasticNet R squared: 0.8442220994403307
Tuned ElasticNet MSE: 10.144762014599413Preprocessing and pipelines
Pipelines, and how scikit-learn allows for transformers and estimators to be chained together and used as a single unit. Preprocessing techniques will be introduced as a way to enhance model performance, and pipelines will tie together concepts from previous sections.
Preprocessing data
Dealing with categorical features
- Scikit-learn will not accept categorical features by default
- Need to encode categorical features numerically
- Convert to ‘dummy variables’
- 0: Observation was NOT that category
- 1: Observation was that category
Dealing with categorical features in Python
- scikit-learn:
OneHotEncoder()- pandas:
get_dummies()
Automobile dataset
- mpg:Target Variable
- Origin:Categorical Feature
autos = pd.read_csv("datasets/autos.csv")
autos.head()| mpg | displ | hp | weight | accel | origin | size | |
|---|---|---|---|---|---|---|---|
| 0 | 18.0 | 250.0 | 88 | 3139 | 14.5 | US | 15.0 |
| 1 | 9.0 | 304.0 | 193 | 4732 | 18.5 | US | 20.0 |
| 2 | 36.1 | 91.0 | 60 | 1800 | 16.4 | Asia | 10.0 |
| 3 | 18.5 | 250.0 | 98 | 3525 | 19.0 | US | 15.0 |
| 4 | 34.3 | 97.0 | 78 | 2188 | 15.8 | Europe | 10.0 |
autos.info()<class 'pandas.core.frame.DataFrame'>
RangeIndex: 392 entries, 0 to 391
Data columns (total 7 columns):
# Column Non-Null Count Dtype
--- ------ -------------- -----
0 mpg 392 non-null float64
1 displ 392 non-null float64
2 hp 392 non-null int64
3 weight 392 non-null int64
4 accel 392 non-null float64
5 origin 392 non-null object
6 size 392 non-null float64
dtypes: float64(4), int64(2), object(1)
memory usage: 21.6+ KBautos.describe()| mpg | displ | hp | weight | accel | size | |
|---|---|---|---|---|---|---|
| count | 392.000000 | 392.000000 | 392.000000 | 392.000000 | 392.000000 | 392.000000 |
| mean | 23.445918 | 194.411990 | 104.469388 | 2977.584184 | 15.541327 | 13.679847 |
| std | 7.805007 | 104.644004 | 38.491160 | 849.402560 | 2.758864 | 4.264458 |
| min | 9.000000 | 68.000000 | 46.000000 | 1613.000000 | 8.000000 | 7.500000 |
| 25% | 17.000000 | 105.000000 | 75.000000 | 2225.250000 | 13.775000 | 10.000000 |
| 50% | 22.750000 | 151.000000 | 93.500000 | 2803.500000 | 15.500000 | 10.000000 |
| 75% | 29.000000 | 275.750000 | 126.000000 | 3614.750000 | 17.025000 | 20.000000 |
| max | 46.600000 | 455.000000 | 230.000000 | 5140.000000 | 24.800000 | 20.000000 |
autos.shape(392, 7)EDA w/ categorical feature
_ = sns.boxplot(data=autos, x="origin", y="mpg", order=['Asia', 'US', 'Europe'])
plt.show()
Encoding dummy variables
autos_origin = pd.get_dummies(autos)
autos_origin.head()| mpg | displ | hp | weight | accel | size | origin_Asia | origin_Europe | origin_US | |
|---|---|---|---|---|---|---|---|---|---|
| 0 | 18.0 | 250.0 | 88 | 3139 | 14.5 | 15.0 | 0 | 0 | 1 |
| 1 | 9.0 | 304.0 | 193 | 4732 | 18.5 | 20.0 | 0 | 0 | 1 |
| 2 | 36.1 | 91.0 | 60 | 1800 | 16.4 | 10.0 | 1 | 0 | 0 |
| 3 | 18.5 | 250.0 | 98 | 3525 | 19.0 | 15.0 | 0 | 0 | 1 |
| 4 | 34.3 | 97.0 | 78 | 2188 | 15.8 | 10.0 | 0 | 1 | 0 |
autos_origin = autos_origin.drop("origin_Asia", axis=1)
autos_origin.head()| mpg | displ | hp | weight | accel | size | origin_Europe | origin_US | |
|---|---|---|---|---|---|---|---|---|
| 0 | 18.0 | 250.0 | 88 | 3139 | 14.5 | 15.0 | 0 | 1 |
| 1 | 9.0 | 304.0 | 193 | 4732 | 18.5 | 20.0 | 0 | 1 |
| 2 | 36.1 | 91.0 | 60 | 1800 | 16.4 | 10.0 | 0 | 0 |
| 3 | 18.5 | 250.0 | 98 | 3525 | 19.0 | 15.0 | 0 | 1 |
| 4 | 34.3 | 97.0 | 78 | 2188 | 15.8 | 10.0 | 1 | 0 |
Linear regression with dummy variables
X_autos_origin = autos_origin[["origin_Europe", "origin_US"]].values
y_autos_origin = autos_origin['mpg'].valuesX_train_autos_origin, X_test_autos_origin, y_train_autos_origin, y_test_autos_origin, = train_test_split(X_autos_origin,
y_autos_origin,
test_size=.3,
random_state=42)
ridge_autos_origin = Ridge(alpha=.5, normalize=True).fit(X_train_autos_origin, y_train_autos_origin)
ridge_autos_origin.score(X_test_autos_origin, y_test_autos_origin)0.3241789154336545Exploring categorical features
The Gapminder dataset that we worked with in previous section also contained a categorical 'Region' feature, which we dropped since we did not have the tools to deal with it. Now however, we do, so we have added it back in!
We will explore this feature. Boxplots are particularly useful for visualizing categorical features such as this.
gapminder.head()| population | fertility | HIV | CO2 | BMI_male | GDP | BMI_female | life | child_mortality | |
|---|---|---|---|---|---|---|---|---|---|
| 0 | 34811059.0 | 2.73 | 0.1 | 3.328945 | 24.59620 | 12314.0 | 129.9049 | 75.3 | 29.5 |
| 1 | 19842251.0 | 6.43 | 2.0 | 1.474353 | 22.25083 | 7103.0 | 130.1247 | 58.3 | 192.0 |
| 2 | 40381860.0 | 2.24 | 0.5 | 4.785170 | 27.50170 | 14646.0 | 118.8915 | 75.5 | 15.4 |
| 3 | 2975029.0 | 1.40 | 0.1 | 1.804106 | 25.35542 | 7383.0 | 132.8108 | 72.5 | 20.0 |
| 4 | 21370348.0 | 1.96 | 0.1 | 18.016313 | 27.56373 | 41312.0 | 117.3755 | 81.5 | 5.2 |
gapminder_2 = pd.read_csv("datasets/gapminder_2.csv")
gapminder_2.head()| population | fertility | HIV | CO2 | BMI_male | GDP | BMI_female | life | child_mortality | Region | |
|---|---|---|---|---|---|---|---|---|---|---|
| 0 | 34811059.0 | 2.73 | 0.1 | 3.328945 | 24.59620 | 12314.0 | 129.9049 | 75.3 | 29.5 | Middle East & North Africa |
| 1 | 19842251.0 | 6.43 | 2.0 | 1.474353 | 22.25083 | 7103.0 | 130.1247 | 58.3 | 192.0 | Sub-Saharan Africa |
| 2 | 40381860.0 | 2.24 | 0.5 | 4.785170 | 27.50170 | 14646.0 | 118.8915 | 75.5 | 15.4 | America |
| 3 | 2975029.0 | 1.40 | 0.1 | 1.804106 | 25.35542 | 7383.0 | 132.8108 | 72.5 | 20.0 | Europe & Central Asia |
| 4 | 21370348.0 | 1.96 | 0.1 | 18.016313 | 27.56373 | 41312.0 | 117.3755 | 81.5 | 5.2 | East Asia & Pacific |
# Create a boxplot of life expectancy per region
gapminder_2.boxplot("life", "Region", rot=60)
# Show the plot
plt.show()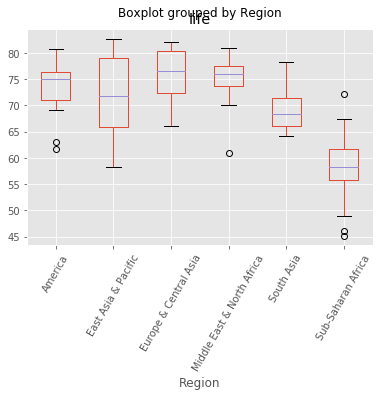
Exploratory data analysis should always be the precursor to model building.
Creating dummy variables
scikit-learn does not accept non-numerical features. The 'Region' feature contains very useful information that can predict life expectancy. For example, Sub-Saharan Africa has a lower life expectancy compared to Europe and Central Asia. Therefore, if we are trying to predict life expectancy, it would be preferable to retain the 'Region' feature. To do this, we need to binarize it by creating dummy variables, which is what we will do.
# Create dummy variables with drop_first=True: df_region
gapminder_region = pd.get_dummies(gapminder_2, drop_first=True)
# Print the new columns of df_region
print(gapminder_region.columns)Index(['population', 'fertility', 'HIV', 'CO2', 'BMI_male', 'GDP',
'BMI_female', 'life', 'child_mortality', 'Region_East Asia & Pacific',
'Region_Europe & Central Asia', 'Region_Middle East & North Africa',
'Region_South Asia', 'Region_Sub-Saharan Africa'],
dtype='object')gapminder_region.head()| population | fertility | HIV | CO2 | BMI_male | GDP | BMI_female | life | child_mortality | Region_East Asia & Pacific | Region_Europe & Central Asia | Region_Middle East & North Africa | Region_South Asia | Region_Sub-Saharan Africa | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 0 | 34811059.0 | 2.73 | 0.1 | 3.328945 | 24.59620 | 12314.0 | 129.9049 | 75.3 | 29.5 | 0 | 0 | 1 | 0 | 0 |
| 1 | 19842251.0 | 6.43 | 2.0 | 1.474353 | 22.25083 | 7103.0 | 130.1247 | 58.3 | 192.0 | 0 | 0 | 0 | 0 | 1 |
| 2 | 40381860.0 | 2.24 | 0.5 | 4.785170 | 27.50170 | 14646.0 | 118.8915 | 75.5 | 15.4 | 0 | 0 | 0 | 0 | 0 |
| 3 | 2975029.0 | 1.40 | 0.1 | 1.804106 | 25.35542 | 7383.0 | 132.8108 | 72.5 | 20.0 | 0 | 1 | 0 | 0 | 0 |
| 4 | 21370348.0 | 1.96 | 0.1 | 18.016313 | 27.56373 | 41312.0 | 117.3755 | 81.5 | 5.2 | 1 | 0 | 0 | 0 | 0 |
Now that we have created the dummy variables, we can use the 'Region' feature to predict life expectancy!
Regression with categorical features
We’ll use ridge regression to perform 5-fold cross-validation.
X_gapminder_region = gapminder_region.drop("life", axis=1).values
y_gapminder_region = gapminder_region.life.values# Instantiate a ridge regressor: ridge
ridge_gapminder_region = Ridge(alpha=.5, normalize=True)
# Perform 5-fold cross-validation: ridge_cv
ridge_cv_gapminder_region = cross_val_score(ridge_gapminder_region, X_gapminder_region, y_gapminder_region, cv=5)
# Print the cross-validated scores
print(ridge_cv_gapminder_region)[0.86808336 0.80623545 0.84004203 0.7754344 0.87503712]We now know how to build models using data that includes categorical features.
Handling missing data
Imputing missing data
- Making an educated guess about the missing values
- Example: Using the mean of the non-missing entries
PIMA Indians dataset
pidd.head()| pregnancies | glucose | diastolic | triceps | insulin | bmi | dpf | age | diabetes | |
|---|---|---|---|---|---|---|---|---|---|
| 0 | 6 | 148 | 72 | 35.00000 | 155.548223 | 33.6 | 0.627 | 50 | 1 |
| 1 | 1 | 85 | 66 | 29.00000 | 155.548223 | 26.6 | 0.351 | 31 | 0 |
| 2 | 8 | 183 | 64 | 29.15342 | 155.548223 | 23.3 | 0.672 | 32 | 1 |
| 3 | 1 | 89 | 66 | 23.00000 | 94.000000 | 28.1 | 0.167 | 21 | 0 |
| 4 | 0 | 137 | 40 | 35.00000 | 168.000000 | 43.1 | 2.288 | 33 | 1 |
pidd.info()<class 'pandas.core.frame.DataFrame'>
RangeIndex: 768 entries, 0 to 767
Data columns (total 9 columns):
# Column Non-Null Count Dtype
--- ------ -------------- -----
0 pregnancies 768 non-null int64
1 glucose 768 non-null int64
2 diastolic 768 non-null int64
3 triceps 768 non-null float64
4 insulin 768 non-null float64
5 bmi 768 non-null float64
6 dpf 768 non-null float64
7 age 768 non-null int64
8 diabetes 768 non-null int64
dtypes: float64(4), int64(5)
memory usage: 54.1 KBpidd.insulin.replace(0, np.nan, inplace=True)
pidd.bmi.replace(0, np.nan, inplace=True)
pidd.triceps.replace(0, np.nan, inplace=True)
pidd.info()<class 'pandas.core.frame.DataFrame'>
RangeIndex: 768 entries, 0 to 767
Data columns (total 9 columns):
# Column Non-Null Count Dtype
--- ------ -------------- -----
0 pregnancies 768 non-null int64
1 glucose 768 non-null int64
2 diastolic 768 non-null int64
3 triceps 768 non-null float64
4 insulin 768 non-null float64
5 bmi 768 non-null float64
6 dpf 768 non-null float64
7 age 768 non-null int64
8 diabetes 768 non-null int64
dtypes: float64(4), int64(5)
memory usage: 54.1 KBpidd.head()| pregnancies | glucose | diastolic | triceps | insulin | bmi | dpf | age | diabetes | |
|---|---|---|---|---|---|---|---|---|---|
| 0 | 6 | 148 | 72 | 35.00000 | 155.548223 | 33.6 | 0.627 | 50 | 1 |
| 1 | 1 | 85 | 66 | 29.00000 | 155.548223 | 26.6 | 0.351 | 31 | 0 |
| 2 | 8 | 183 | 64 | 29.15342 | 155.548223 | 23.3 | 0.672 | 32 | 1 |
| 3 | 1 | 89 | 66 | 23.00000 | 94.000000 | 28.1 | 0.167 | 21 | 0 |
| 4 | 0 | 137 | 40 | 35.00000 | 168.000000 | 43.1 | 2.288 | 33 | 1 |
Dropping missing data
The voting dataset1 contained a bunch of missing values that we dealt with for you behind the scenes.
votes2 = pd.read_csv("datasets/votes2.csv")
votes2.head()| party | infants | water | budget | physician | salvador | religious | satellite | aid | missile | immigration | synfuels | education | superfund | crime | duty_free_exports | eaa_rsa | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 0 | republican | 0 | 1 | 0 | 1 | 1 | 1 | 0 | 0 | 0 | 1 | ? | 1 | 1 | 1 | 0 | 1 |
| 1 | republican | 0 | 1 | 0 | 1 | 1 | 1 | 0 | 0 | 0 | 0 | 0 | 1 | 1 | 1 | 0 | ? |
| 2 | democrat | ? | 1 | 1 | ? | 1 | 1 | 0 | 0 | 0 | 0 | 1 | 0 | 1 | 1 | 0 | 0 |
| 3 | democrat | 0 | 1 | 1 | 0 | ? | 1 | 0 | 0 | 0 | 0 | 1 | 0 | 1 | 0 | 0 | 1 |
| 4 | democrat | 1 | 1 | 1 | 0 | 1 | 1 | 0 | 0 | 0 | 0 | 1 | ? | 1 | 1 | 1 | 1 |
there are certain data points labeled with a '?'. These denote missing values. We will convert the '?'s to NaNs, and then drop the rows that contain them from the DataFrame.
# Convert '?' to NaN
votes2[votes2 == "?"] = np.nan
# Print the number of NaNs
display(votes2.isnull().sum())
# Print shape of original DataFrame
print("Shape of Original DataFrame: {}".format(votes2.shape))
# Print shape of new DataFrame
print("Shape of DataFrame After Dropping All Rows with Missing Values: {}".format(votes2.dropna().shape))party 0
infants 12
water 48
budget 11
physician 11
salvador 15
religious 11
satellite 14
aid 15
missile 22
immigration 7
synfuels 21
education 31
superfund 25
crime 17
duty_free_exports 28
eaa_rsa 104
dtype: int64Shape of Original DataFrame: (435, 17)
Shape of DataFrame After Dropping All Rows with Missing Values: (232, 17)When many values in a dataset are missing, if you drop them, you may end up throwing away valuable information along with the missing data. It’s better instead to develop an imputation strategy. This is where domain knowledge is useful, but in the absence of it, you can impute missing values with the mean or the median of the row or column that the missing value is in.
Imputing missing data in a ML Pipeline I
there are many steps to building a model, from creating training and test sets, to fitting a classifier or regressor, to tuning its parameters, to evaluating its performance on new data. Imputation can be seen as the first step of this machine learning process, the entirety of which can be viewed within the context of a pipeline. Scikit-learn provides a pipeline constructor that allows you to piece together these steps into one process and thereby simplify your workflow.
We will be setting up a pipeline with two steps: the imputation step, followed by the instantiation of a classifier. We’ve seen three classifiers in this course so far: k-NN, logistic regression, and the decision tree. Here we will be using the SVM (Support Vector Machine)
votes2.head()| party | infants | water | budget | physician | salvador | religious | satellite | aid | missile | immigration | synfuels | education | superfund | crime | duty_free_exports | eaa_rsa | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 0 | republican | 0 | 1 | 0 | 1 | 1 | 1 | 0 | 0 | 0 | 1 | NaN | 1 | 1 | 1 | 0 | 1 |
| 1 | republican | 0 | 1 | 0 | 1 | 1 | 1 | 0 | 0 | 0 | 0 | 0 | 1 | 1 | 1 | 0 | NaN |
| 2 | democrat | NaN | 1 | 1 | NaN | 1 | 1 | 0 | 0 | 0 | 0 | 1 | 0 | 1 | 1 | 0 | 0 |
| 3 | democrat | 0 | 1 | 1 | 0 | NaN | 1 | 0 | 0 | 0 | 0 | 1 | 0 | 1 | 0 | 0 | 1 |
| 4 | democrat | 1 | 1 | 1 | 0 | 1 | 1 | 0 | 0 | 0 | 0 | 1 | NaN | 1 | 1 | 1 | 1 |
votes2.info()<class 'pandas.core.frame.DataFrame'>
RangeIndex: 435 entries, 0 to 434
Data columns (total 17 columns):
# Column Non-Null Count Dtype
--- ------ -------------- -----
0 party 435 non-null object
1 infants 423 non-null object
2 water 387 non-null object
3 budget 424 non-null object
4 physician 424 non-null object
5 salvador 420 non-null object
6 religious 424 non-null object
7 satellite 421 non-null object
8 aid 420 non-null object
9 missile 413 non-null object
10 immigration 428 non-null object
11 synfuels 414 non-null object
12 education 404 non-null object
13 superfund 410 non-null object
14 crime 418 non-null object
15 duty_free_exports 407 non-null object
16 eaa_rsa 331 non-null object
dtypes: object(17)
memory usage: 57.9+ KB# Setup the Imputation transformer: imp
imp_votes = SimpleImputer(missing_values=np.nan, strategy="most_frequent")
# Instantiate the SVC classifier: clf
clf_votes = SVC()
# Setup the pipeline with the required steps: steps
steps_votes = [('imputation', imp_votes),
('SVM', clf_votes)]Having set up the pipeline steps, we can now use it for classification.
Imputing missing data in a ML Pipeline II
Having setup the steps of the pipeline we will now use it on the voting dataset to classify a Congressman’s party affiliation. What makes pipelines so incredibly useful is the simple interface that they provide.
X_votes[:5]array([[0, 1, 0, 1, 1, 1, 0, 0, 0, 1, 0, 1, 1, 1, 0, 1],
[0, 1, 0, 1, 1, 1, 0, 0, 0, 0, 0, 1, 1, 1, 0, 1],
[0, 1, 1, 0, 1, 1, 0, 0, 0, 0, 1, 0, 1, 1, 0, 0],
[0, 1, 1, 0, 1, 1, 0, 0, 0, 0, 1, 0, 1, 0, 0, 1],
[1, 1, 1, 0, 1, 1, 0, 0, 0, 0, 1, 0, 1, 1, 1, 1]], dtype=int64)votes.head()| party | infants | water | budget | physician | salvador | religious | satellite | aid | missile | immigration | synfuels | education | superfund | crime | duty_free_exports | eaa_rsa | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 0 | republican | 0 | 1 | 0 | 1 | 1 | 1 | 0 | 0 | 0 | 1 | 0 | 1 | 1 | 1 | 0 | 1 |
| 1 | republican | 0 | 1 | 0 | 1 | 1 | 1 | 0 | 0 | 0 | 0 | 0 | 1 | 1 | 1 | 0 | 1 |
| 2 | democrat | 0 | 1 | 1 | 0 | 1 | 1 | 0 | 0 | 0 | 0 | 1 | 0 | 1 | 1 | 0 | 0 |
| 3 | democrat | 0 | 1 | 1 | 0 | 1 | 1 | 0 | 0 | 0 | 0 | 1 | 0 | 1 | 0 | 0 | 1 |
| 4 | democrat | 1 | 1 | 1 | 0 | 1 | 1 | 0 | 0 | 0 | 0 | 1 | 0 | 1 | 1 | 1 | 1 |
X_votes = votes.drop("party", axis=1)
y_votes = votes.partyX_train_votes, X_test_votes, y_train_votes, y_test_votes = train_test_split(X_votes, y_votes, test_size=.3, random_state=42)# Create the pipeline: pipeline
pipeline_votes = Pipeline(steps_votes)
# Fit the pipeline to the train set
pipeline_votes.fit(X_train_votes, y_train_votes)
# Predict the labels of the test set
y_pred_votes = pipeline_votes.predict(X_test_votes)
# Compute metrics
print(classification_report(y_test_votes, y_pred_votes)) precision recall f1-score support
democrat 0.98 0.96 0.97 85
republican 0.94 0.96 0.95 46
accuracy 0.96 131
macro avg 0.96 0.96 0.96 131
weighted avg 0.96 0.96 0.96 131
Centering and scaling
Why scale your data?
- Many models use some form of distance to inform them
- Features on larger scales can unduly influence the model
- Example: k-NN uses distance explicitly when making predictions
- We want features to be on a similar scale
- Normalizing (or scaling and centering)
Ways to normalize your data
- Standardization: Subtract the mean and divide by variance
- All features are centered around zero and have variance one
- Can also subtract the minimum and divide by the range
- Minimum zero and maximum one
- Can also normalize so the data ranges from -1 to +1
Centering and scaling your data
the performance of a model can improve if the features are scaled. Note that this is not always the case: In the Congressional voting records dataset, for example, all of the features are binary. In such a situation, scaling will have minimal impact. We will explore scalling on White Wine Quality.
wwq = pd.read_csv("datasets/white_wine_quality.csv")
wwq.head()| fixed acidity | volatile acidity | citric acid | residual sugar | chlorides | free sulfur dioxide | total sulfur dioxide | density | pH | sulphates | alcohol | quality | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 0 | 7.0 | 0.27 | 0.36 | 20.7 | 0.045 | 45.0 | 170.0 | 1.0010 | 3.00 | 0.45 | 8.8 | 6 |
| 1 | 6.3 | 0.30 | 0.34 | 1.6 | 0.049 | 14.0 | 132.0 | 0.9940 | 3.30 | 0.49 | 9.5 | 6 |
| 2 | 8.1 | 0.28 | 0.40 | 6.9 | 0.050 | 30.0 | 97.0 | 0.9951 | 3.26 | 0.44 | 10.1 | 6 |
| 3 | 7.2 | 0.23 | 0.32 | 8.5 | 0.058 | 47.0 | 186.0 | 0.9956 | 3.19 | 0.40 | 9.9 | 6 |
| 4 | 7.2 | 0.23 | 0.32 | 8.5 | 0.058 | 47.0 | 186.0 | 0.9956 | 3.19 | 0.40 | 9.9 | 6 |
X_wwq = pd.read_csv("datasets/X_wwq.csv").values
X_wwq[:5]array([[7.000e+00, 2.700e-01, 3.600e-01, 2.070e+01, 4.500e-02, 4.500e+01,
1.700e+02, 1.001e+00, 3.000e+00, 4.500e-01, 8.800e+00],
[6.300e+00, 3.000e-01, 3.400e-01, 1.600e+00, 4.900e-02, 1.400e+01,
1.320e+02, 9.940e-01, 3.300e+00, 4.900e-01, 9.500e+00],
[8.100e+00, 2.800e-01, 4.000e-01, 6.900e+00, 5.000e-02, 3.000e+01,
9.700e+01, 9.951e-01, 3.260e+00, 4.400e-01, 1.010e+01],
[7.200e+00, 2.300e-01, 3.200e-01, 8.500e+00, 5.800e-02, 4.700e+01,
1.860e+02, 9.956e-01, 3.190e+00, 4.000e-01, 9.900e+00],
[7.200e+00, 2.300e-01, 3.200e-01, 8.500e+00, 5.800e-02, 4.700e+01,
1.860e+02, 9.956e-01, 3.190e+00, 4.000e-01, 9.900e+00]])y_wwq = np.array([False, False, False, False, False, False, False, False, False,
False, True, True, True, False, True, False, False, False,
False, True, False, False, False, True, False, False, False,
False, False, False, False, False, False, False, True, True,
True, False, True, True, False, False, False, False, False,
False, True, True, False, True, False, False, False, False,
False, False, False, False, False, False, False, False, True,
False, False, True, False, True, False, True, False, True,
True, False, False, True, False, False, True, True, False,
False, True, False, True, False, False, False, True, False,
False, True, False, False, False, False, False, False, True,
False, True, True, True, True, True, False, True, False,
False, True, False, True, True, True, True, True, False,
False, True, True, True, True, True, False, False, False,
True, False, False, False, True, False, True, True, True,
True, False, True, False, False, True, True, False, False,
False, False, False, True, False, False, False, False, False,
True, False, False, False, False, False, False, False, True,
True, False, True, True, False, False, True, True, False,
False, True, False, True, False, True, True, True, False,
False, True, True, False, True, True, False, True, False,
True, False, True, False, True, True, False, True, True,
True, True, True, True, True, False, True, True, True,
True, True, False, True, False, True, False, False, True,
True, True, True, True, True, False, False, False, False,
True, False, False, False, True, True, False, False, False,
False, False, False, False, False, False, True, True, False,
False, True, False, False, False, False, True, True, True,
True, True, False, False, False, False, False, True, False,
True, True, False, False, True, False, True, False, False,
False, True, True, True, True, False, False, True, True,
False, False, False, True, True, True, True, False, False,
False, False, False, False, True, False, True, False, True,
False, False, False, False, False, False, False, False, False,
True, False, False, False, False, False, False, False, True,
False, False, True, False, False, False, True, False, False,
True, True, False, False, False, True, False, True, False,
True, True, False, False, False, True, False, False, False,
False, True, False, False, False, False, False, True, False,
False, False, False, False, False, False, False, False, False,
False, True, False, False, False, False, False, False, False,
True, False, False, True, False, False, False, False, False,
False, False, False, False, False, False, False, False, False,
True, False, False, False, True, False, False, True, True,
True, False, True, False, False, True, True, True, False,
True, False, True, False, True, False, False, True, True,
False, False, False, True, False, False, False, False, False,
False, False, False, False, True, True, True, True, True,
False, True, False, False, True, False, False, True, False,
False, False, False, False, True, True, False, False, False,
True, True, False, False, False, False, False, True, False,
True, True, True, True, False, True, True, False, False,
True, True, False, True, False, False, False, True, False,
False, False, False, True, False, True, True, True, False,
False, False, False, False, False, False, False, False, False,
False, True, False, True, True, False, False, False, True,
False, False, True, False, False, False, False, False, False,
False, False, False, True, False, False, True, True, True,
False, False, True, False, True, False, False, False, False,
True, False, False, False, True, True, False, True, False,
True, True, False, False, False, False, False, False, False,
True, False, False, False, False, False, False, True, False,
True, False, False, True, False, False, True, False, False,
True, False, False, True, False, True, False, False, False,
False, False, False, False, True, True, False, False, False,
False, False, False, False, False, True, False, True, True,
True, False, True, False, False, False, False, False, True,
True, False, False, True, True, True, False, False, False,
True, True, True, True, False, False, False, False, True,
True, False, True, True, False, True, False, False, False,
True, True, False, True, False, False, False, True, True,
True, False, True, False, True, True, True, True, False,
True, False, False, False, False, False, False, False, False,
True, True, True, True, False, True, True, False, True,
False, False, False, True, False, False, False, False, True,
False, False, False, False, False, True, True, False, True,
True, True, False, True, False, False, True, False, True,
True, False, True, False, True, True, True, True, False,
True, False, False, True, True, False, False, True, True,
False, False, True, False, False, False, True, False, False,
True, True, False, False, False, True, False, True, True,
True, False, False, False, False, True, False, False, True,
False, False, True, False, False, True, True, False, False,
False, False, False, False, False, False, False, True, False,
True, False, False, False, False, False, True, False, False,
False, True, False, False, False, False, False, False, True,
False, False, False, True, False, False, True, False, False,
False, True, False, False, False, False, False, False, False,
True, False, True, True, False, False, False, False, False,
True, True, False, False, False, True, False, False, False,
True, False, False, False, False, True, True, True, True,
True, False, False, True, False, True, False, False, False,
False, False, False, True, False, False, False, False, False,
False, False, False, False, False, False, False, False, False,
True, True, False, False, False, True, False, True, False,
False, True, True, False, True, False, False, False, False,
True, False, False, False, False, False, True, True, False,
True, True, False, False, False, False, False, False, False,
False, False, False, False, False, False, False, False, False,
False, True, False, False, False, False, False, False, False,
False, False, True, False, False, True, True, False, True,
True, True, True, True, True, True, True, False, False,
False, False, False, False, False, False, False, True, True,
False, True, True, False, True, False, True, True, True,
True, True, False, False, True, False, False, False, False,
False, True, True, True, True, True, False, False, True,
False, True, True, False, False, False, False, False, False,
False, False, True, False, False, False, False, False, False,
False, False, False, True, False, True, False, True, False,
False, False, False, True, False, False, False, True, False,
False, True, True, True, False, False, True, False, False,
False, False, True, False, False, False, False, False, True,
False, False, False, False, False, True, True, True, True,
False, False, False, False, False, False, False, False, False,
False, True, False, True, False, False, False, False, True,
False, True, True, False, True, True, True, True, True,
False, True, True, True, True, False, True, False, True,
True, True, True, False, True, False, True, False, True,
True, True, False, True, False, False, False, False, False,
True, False, True, False, False, False, True, True, False,
False, False, False, False, False, True, False, False, False,
False, False, False, False, False, True, False, False, False,
False, False, False, False, True, True, True, False, False,
False, False, True, False, False, False, True, True, False,
True, True, True, True, False, False, False, True, True,
False, False, False, True, False, False, False, True, True,
True, False, False, False, True, True, False, True, True,
True, True, True, True, True, False, True, True, True,
True, False, True, True, False, False, False, False, False,
False, False, False, False, True, True, False, True, False,
False, True, False, True, False, True, False, False, True,
True, True, False, True, False, False, True, True, False,
False, True, False, False, False, False, False, False, False,
False, False, False, False, False, True, False, False, True,
False, True, False, False, False, True, False, False, False,
True, False, True, False, False, False, False, False, True,
False, False, False, True, False, True, False, False, False,
False, False, True, False, True, False, True, False, False,
False, False, True, True, False, False, False, True, False,
True, False, False, True, False, False, False, False, False,
False, False, False, False, True, False, False, False, True,
False, False, True, True, True, False, False, False, True,
False, False, False, False, True, False, False, False, False,
False, False, False, False, False, False, True, True, False,
False, False, False, True, False, True, False, False, True,
False, False, True, False, False, False, False, True, True,
False, False, False, True, False, True, False, False, False,
True, False, False, False, False, False, True, True, False,
False, False, False, False, False, True, True, False, False,
True, False, False, False, False, False, False, False, True,
False, False, False, True, True, True, False, False, False,
False, False, False, False, True, True, False, False, True,
False, True, False, False, False, False, False, False, False,
False, False, False, False, False, False, False, False, True,
False, False, False, False, False, False, False, False, False,
False, False, False, False, False, False, False, False, False,
True, True, False, False, False, False, False, False, False,
False, False, True, False, True, True, False, True, False,
True, True, False, False, False, True, False, True, True,
True, False, False, True, True, True, False, True, True,
True, True, True, False, False, False, False, False, False,
False, False, True, False, False, True, True, False, False,
False, False, False, False, False, False, False, True, False,
False, True, True, False, False, True, False, True, False,
True, False, False, False, False, False, False, True, True,
False, False, False, False, True, False, True, True, False,
False, False, True, False, False, False, False, True, True,
False, False, False, False, False, False, True, True, False,
False, False, False, False, False, False, False, True, False,
True, True, False, False, True, False, False, False, False,
True, False, False, False, False, False, True, False, False,
False, False, True, False, False, True, False, False, False,
False, False, False, False, False, False, False, False, False,
False, True, True, False, True, False, False, True, False,
True, True, False, True, True, False, False, True, True,
False, False, True, True, True, True, False, False, True,
False, True, True, False, True, False, False, False, False,
False, False, False, True, False, False, False, False, False,
True, False, True, False, False, False, True, False, False,
False, False, False, True, False, False, False, False, False,
False, True, False, False, False, False, False, True, False,
False, True, False, False, False, False, False, True, False,
True, True, True, False, False, False, False, False, False,
False, True, True, False, False, True, False, True, True,
True, False, True, True, False, False, False, False, True,
False, True, False, False, False, False, False, True, True,
False, True, False, False, False, True, True, False, False,
False, False, False, True, False, True, False, True, False,
False, True, False, False, False, False, False, False, False,
True, True, True, True, False, False, True, True, True,
True, True, False, False, True, False, False, False, True,
False, False, False, False, False, False, False, False, False,
True, False, False, False, True, False, False, False, True,
False, True, True, False, False, False, False, True, True,
True, False, True, False, False, False, False, False, False,
False, True, True, True, True, False, False, False, False,
True, True, True, True, True, False, False, False, False,
False, False, False, True, False, False, False, True, True,
False, False, False, False, False, False, True, False, True,
True, False, True, True, False, True, False, False, False,
False, True, False, False, False, True, True, False, True,
False, False, True, False, False, False, False, False, True,
False, False, False, True, False, False, True, True, True,
False, False, False, False, False, False, False, False, True,
False, False, False, False, True, False, False, False, False,
True, True, True, True, False, False, False, False, True,
False, False, True, True, True, False, False, False, False,
True, False, False, False, False, False, False, False, False,
True, True, True, True, False, False, False, False, True,
False, True, True, True, False, True, True, True, True,
True, True, True, True, True, False, False, False, False,
False, False, True, False, True, False, False, True, False,
False, True, True, False, False, False, False, False, True,
False, False, False, False, True, True, True, True, False,
False, True, False, True, False, True, False, True, False,
True, False, False, True, True, True, True, True, True,
True, False, False, True, False, True, True, True, False,
False, True, True, True, False, False, True, True, True,
True, False, True, False, False, False, False, False, False,
False, True, True, True, False, True, False, False, False,
False, False, False, False, False, False, False, True, False,
False, True, False, True, False, False, False, False, False,
False, False, True, True, False, False, False, False, False,
False, True, False, False, True, True, False, True, False,
False, True, False, False, False, False, False, False, True,
True, True, True, False, False, False, True, False, True,
True, True, False, True, True, True, False, True, False,
False, False, True, True, True, True, False, True, True,
True, True, True, False, True, False, True, True, True,
False, True, True, False, True, True, True, False, False,
False, False, True, True, False, True, True, False, False,
True, True, True, False, True, False, False, True, False,
False, False, True, True, True, True, True, True, True,
True, False, True, True, False, True, True, True, True,
True, False, False, False, True, True, True, True, False,
True, True, True, False, True, True, False, True, True,
False, True, True, True, False, True, True, False, False,
False, False, True, False, True, True, True, True, True,
True, False, False, False, False, True, True, False, False,
True, False, False, True, True, True, False, False, True,
False, False, False, False, True, True, True, False, False,
False, False, False, True, True, False, False, True, False,
True, False, False, False, True, True, False, False, True,
True, False, False, False, True, False, False, False, True,
False, False, True, False, True, False, True, True, True,
True, False, False, True, False, False, False, False, False,
True, False, True, False, False, False, False, False, False,
False, False, True, True, True, False, False, False, False,
False, True, True, False, True, True, False, False, True,
True, True, True, True, False, True, False, False, True,
False, False, True, True, False, False, True, False, True,
False, True, False, False, True, False, True, True, False,
False, False, False, False, False, True, False, True, False,
False, False, True, False, False, True, True, True, True,
True, True, True, False, False, False, False, False, False,
False, False, False, False, False, False, True, True, True,
False, False, True, True, True, False, True, False, True,
False, False, False, False, True, True, False, False, True,
False, False, True, False, True, False, True, True, False,
True, False, False, True, True, True, True, True, False,
False, True, False, True, False, False, True, False, False,
True, True, False, False, False, False, False, False, True,
True, True, False, True, False, True, True, True, True,
False, False, False, False, True, True, True, False, False,
False, False, True, True, True, False, False, False, False,
True, True, True, True, False, False, False, False, False,
True, True, True, False, False, False, True, True, False,
False, True, True, False, False, True, True, False, False,
True, True, True, True, False, True, True, True, False,
True, True, False, True, False, False, False, False, False,
True, True, True, False, False, True, True, False, True,
False, True, False, True, True, True, True, True, False,
False, False, True, True, False, False, True, False, False,
False, False, True, True, True, True, False, True, True,
True, False, False, True, True, False, False, False, False,
True, False, False, False, False, True, False, False, False,
True, False, True, False, False, False, False, True, False,
True, True, True, True, True, True, False, True, True,
True, True, False, False, False, False, False, True, False,
False, False, True, False, False, False, False, False, True,
True, False, False, True, True, False, False, False, False,
True, False, True, True, True, False, False, False, False,
False, True, True, False, True, False, False, False, True,
True, False, True, True, False, True, False, False, False,
True, False, False, True, False, True, False, True, False,
True, False, True, False, False, True, False, True, True,
True, False, True, True, False, False, True, False, False,
False, False, False, False, False, False, True, False, False,
True, True, False, True, True, False, True, False, False,
True, False, False, False, False, False, False, False, False,
True, True, False, True, True, False, False, False, False,
False, True, True, False, False, True, True, False, False,
False, True, False, False, True, True, True, False, True,
True, False, False, False, True, True, True, False, False,
False, True, False, False, True, True, False, True, True,
True, True, False, False, False, False, False, False, False,
False, True, False, False, False, True, False, False, False,
False, False, False, False, False, True, False, False, False,
False, True, False, False, False, True, False, False, False,
False, False, False, False, False, False, True, False, False,
False, True, True, True, False, False, False, True, False,
True, True, True, True, False, False, False, True, False,
False, False, False, True, False, False, False, True, False,
False, False, False, True, True, True, False, False, False,
False, False, True, False, False, True, False, False, False,
False, False, False, False, False, False, False, False, False,
True, True, False, True, False, False, False, True, True,
False, False, False, False, False, False, True, False, False,
False, False, False, False, False, False, False, True, True,
False, False, False, False, True, False, True, True, True,
False, True, True, False, True, False, False, True, False,
False, True, True, False, False, False, False, True, True,
False, False, False, True, False, False, False, True, True,
False, True, False, True, True, True, False, False, False,
False, False, False, True, True, True, False, False, False,
False, False, False, False, False, False, False, False, False,
True, False, False, True, True, True, True, True, True,
False, False, False, False, False, False, False, False, False,
False, False, False, False, False, False, True, False, False,
False, False, False, False, False, True, True, False, False,
True, False, True, False, False, False, True, True, True,
False, False, False, True, False, False, True, True, True,
True, True, False, False, True, False, False, True, True,
False, False, False, True, True, False, True, False, False,
False, False, False, True, True, False, False, False, False,
True, False, False, False, False, False, False, False, False,
True, True, True, True, False, False, False, True, True,
False, False, False, False, False, False, False, True, True,
True, True, False, True, False, False, True, False, False,
False, True, False, True, False, False, False, True, False,
True, True, False, False, False, False, False, False, False,
False, False, False, False, False, False, False, False, False,
False, False, False, False, False, False, False, False, False,
False, False, False, False, False, False, False, False, False,
False, False, False, False, True, False, False, False, False,
False, False, False, False, False, False, True, True, False,
False, False, False, True, False, False, False, False, True,
False, False, False, True, False, False, False, True, False,
False, False, False, False, False, True, False, True, True,
True, False, True, False, False, True, True, True, False,
True, True, False, True, True, False, True, True, False,
False, False, False, False, True, True, False, True, True,
True, True, False, True, False, False, False, False, False,
True, False, False, False, False, False, False, False, False,
False, True, False, False, False, False, False, False, True,
False, False, False, True, True, False, False, True, False,
False, False, False, False, False, True, False, True, False,
True, False, True, False, True, False, False, False, True,
False, False, False, False, True, False, False, False, False,
False, False, True, False, False, False, False, False, False,
False, False, False, False, False, False, True, False, False,
False, False, True, False, False, False, False, False, False,
False, False, False, False, False, False, False, False, False,
True, False, True, False, False, True, False, False, False,
True, False, False, True, False, False, False, True, True,
False, False, False, False, False, False, False, False, False,
True, False, False, False, True, True, True, False, True,
False, False, False, True, True, True, False, True, False,
False, False, False, False, False, False, False, False, False,
False, True, False, False, False, False, False, True, False,
False, False, False, False, False, True, False, False, False,
False, False, False, False, False, True, False, False, False,
False, False, False, True, False, False, False, True, False,
False, False, True, True, False, False, False, True, True,
False, False, True, False, True, False, False, False, False,
False, False, False, False, True, True, False, True, False,
False, True, True, True, False, False, True, False, False,
False, False, False, False, True, False, False, False, False,
False, False, False, False, False, False, False, True, True,
False, False, False, False, False, True, False, True, False,
False, False, False, False, False, False, False, False, False,
False, False, False, True, False, False, False, False, False,
False, False, False, False, False, False, False, False, False,
False, False, False, False, False, False, False, False, False,
False, False, False, False, False, False, False, False, True,
False, False, False, False, True, False, False, False, False,
False, False, False, True, False, False, False, False, False,
False, True, True, False, False, True, False, False, True,
False, True, False, False, False, True, False, True, False,
False, False, False, False, False, False, True, False, False,
True, False, False, True, False, False, False, False, False,
True, False, False, True, False, False, False, False, True,
True, True, True, True, False, False, True, False, False,
False, False, False, False, False, True, False, True, False,
True, False, False, False, False, False, False, False, True,
False, False, True, False, False, False, False, False, False,
False, False, False, False, False, False, False, True, False,
False, False, False, False, False, False, False, False, False,
False, False, False, False, False, False, False, False, False,
False, True, False, False, False, False, False, True, False,
False, False, False, False, False, False, False, False, True,
False, True, False, True, True, False, False, False, False,
True, False, True, True, False, False, True, True, False,
True, False, False, False, False, False, False, False, False,
False, False, False, False, False, False, False, False, False,
False, False, False, False, True, False, True, False, True,
False, False, False, False, False, False, False, True, False,
False, False, False, True, False, True, False, True, False,
False, False, False, False, False, False, False, False, False,
True, False, False, False, False, False, False, False, False,
False, False, False, False, False, True, True, False, False,
False, True, False, False, False, True, False, True, True,
False, False, False, False, False, False, False, True, True,
True, False, False, False, False, False, False, False, False,
False, False, False, False, False, False, False, False, False,
False, False, False, True, False, True, False, False, False,
False, False, False, False, False, False, False, False, True,
False, False, False, False, False, True, False, False, False,
False, False, True, False, False, False, True, False, False,
False, False, False, False, False, True, True, False, False,
False, False, False, False, True, False, False, False, False,
True, True, True, False, False, False, False, False, False,
True, True, True, False, True, True, True, False, True,
False, True, True, False, True, True, False, False, False,
False, False, False, True, False, False, True, False, True,
False, True, True, False, False, False, False, False, False,
False, False, True, False, True, False, False, True, True,
True, False, False, True, True, True, False, False, False,
True, True, False, True, True, True, False, True, True,
False, False, True, True, False, True, False, False, False,
True, False, False, True, False, False, True, True, True,
False, True, True, False, False, True, False, False, True,
True, True, True, False, True, False, True, False, True,
True, False, False, True, True, True, True, True, False,
True, False, False, False, True, False, False, True, False,
False, False, True, False, False, False, True, True, False,
True, True, False, False, True, True, False, False, False,
False, False, True, False, True, False, True, False, False,
False, False, False, False, False, False, False, True, True,
False, False, False, False, False, False, True, False, False,
False, False, True, True, True, False, True, True, True,
True, False, False, True, False, False, False, False, False,
False, False, False, True, True, True, False, False, False,
False, False, True, True, True, False, False, False, False,
False, False, False, False, False, False, False, False, False,
False, False, True, False, False, False, True, False, True,
True, True, False, True, True, True, False, False, False,
True, False, True, False, True, False, False, True, True,
False, False, False, False, False, False, True, True, True,
True, False, False, True, False, True, True, False, False,
False, True, False, False, True, True, False, True, False,
False, False, False, False, True, False, False, False, False,
False, False, True, False, False, False, False, False, False,
False, False, False, False, True, False, False, False, True,
False, False, False, False, False, True, False, False, False,
False, False, True, False, False, False, False, False, False,
False, True, True, False, False, True, True, True, True,
False, False, False, False, True, False, False, True, True,
True, False, True, False, True, True, True, True, True,
True, False, False, False, False, False, False, False, False,
False, False, True, False, False, False, True, False, False,
False, True, True, False, True, True, False, False, True,
False, False, True, False, False, True, True, False, True,
False, False, False, False, False, False, True, False, False,
False, False, False, False, False, False, False, False, False,
False, True, False, True, False, False, True, True, False,
False, False, True, False, False, False, True, False, False,
False, False, False, False, False, False, False, True, False,
False, False, False, True, True, True, True, False, False,
True, True, False, False, True, False, True, False, False,
False, False, False, False, False, False, False, False, False,
False, True, False, False, True, True, True, True, False,
False, False, False, False, False, True, True, False, False,
False, False, False, False, True, False, True, False, False,
False, False, False, True, False, False, False, False, False,
True, True, False, False, True, False, True, False, False,
False, False, False, True, True, False, False, False, False,
True, False, False, False, False, False, False, False, False,
True, True, True, True, True, True, False, True, False,
False, True, True, False, True, False, False, False, False,
False, False, False, False, False, True, True, True, True,
False, False, False, False, False, True, True, False, False,
False, False, False, False, False, True, True, True, True,
True, False, False, False, False, False, False, False, False,
False, False, False, True, False, True, False, False, False,
False, False, True, True, False, False, True, True, False,
False, False, False, True, False, False, True, True, False,
True, True, True, False, False, False, False, True, True,
False, False, False, False, False, False, False, False, False,
True, False, False, False, False, True, False, False, False,
True, False, False, False, False, True, True, True, True,
True, True, True, False, False, False, False, False, False,
False, False, False, False, False, False, False, False, False,
False, False, True, False, False, True, False, False, False,
True, False, True, False, True, False, False, True, False,
True, False, True, False, False, True, True, False, False,
True, False, True, False, True, True, False, False, False,
False, True, True, False, False, False, True, True, True,
True, False, False, True, True, True, True, True, True,
False, False, True, True, True, False, False, True, False,
True, False, False, False, False, True, True, False, False,
False, False, False, False, False, False, False, False, True,
False, False, True, False, False, True, False, False, False,
False, True, True, True, False, True, True, False, False,
False, False, True, False, False, True, True, False, False,
True, True, True, False, True, False, False, False, False,
True, True, True, True, False, False, False, False, True,
True, False, False, False, True, True, True, True, True,
False, True, True, False, False, False, False, False, False,
False, False, False, False, False, False, True, True, False,
False, False, False, False, True, True, False, True, True,
True, False, False, False, False, True, False, False, False,
False, False, True, True, False, True, False, False, False,
False, False, False, False, False, False, True, False, False,
False, True, False, True, False, False, True, True, False,
False, False, False, True, True, False, False, False, False,
False, True, True, False, False, False, False, False, False,
False, False, True, True, True, False, False, False, False,
False, False, False, False, True, False, False, False, False,
False, False, False, False, False, False, False, True, False,
False, False, False, False, False, False, False, False, False,
False, False, False, False, False, False, False, True, False,
False, True, False, True, True, True, True, False, True,
False, False, True, False, False, False, False, False, True,
False, False, False, False, False, False, False, False, False,
False, True, False, True, False, False, False, False, False,
False, False, False, False, True, True, True, False, True,
False, False, False, False, False, False, False, False, True,
False, False, False, False, True, False, False, True, False,
True, True, False, False, False, False, False, False, True,
True, False, False, False, False, True, True, False, False,
False, False, False, True, False, False, False, False, False,
False, True, True, False, False, False, False, False, False,
False, False, False, False, False, False, False, False, False,
False, False, False, False, True, False, False, False, False,
False, False, False, False, True, True, False, False, False,
False, False, False, False, False, False, True, False, False,
True, True, True, False, False, True, False, True, False,
False, False, False, False, False, False, False, False, False,
False, False, False, False, False, False, False, False, False,
False, False, True, False, True, False, False, False, False,
True, True, True, False, False, True, False, False, False,
True, False, False, False, False, False, True, False, False,
False, True, False, False, False, False, False, True, False,
False, False, True, False, True, False, False, True, False,
False, False, False, False, False, True, False, False, False,
True, False, False, False, True, False, False, False, True,
False, False, False, True, False, False, True, False, False,
False, True, False, True, False, False, False, True, False,
True, True, True, False, False, False, False, True, False,
False, False, False, False, True, False, False, False, False,
False, False, False, True, False, False, False, False, True,
True, False, False, False, True, False, True, False, False,
False, True, False, False, False, True, False, True, False,
False, False])y_wwq = pd.read_csv("datasets/y_wwq.csv").values
y_wwq[:5]array([[False],
[False],
[False],
[False],
[False]])some features seem to have different units of measurement. 'density', for instance, takes values between 0.98 and 1.04, while 'total sulfur dioxide' ranges from 9 to 440. As a result, it may be worth scaling the features here. We will scale the features and compute the mean and standard deviation of the unscaled features compared to the scaled features.
# Scale the features: X_scaled
X_scaled_wwq = scale(X_wwq)
# Print the mean and standard deviation of the unscaled features
print("Mean of Unscaled Features: {}".format(np.mean(X_wwq)))
print("Standard Deviation of Unscaled Features: {}".format(np.std(X_wwq)))Mean of Unscaled Features: 18.43268707245592
Standard Deviation of Unscaled Features: 41.544947640946354# Print the mean and standard deviation of the scaled features
print("Mean of Scaled Features: {}".format(np.mean(X_scaled_wwq)))
print("Standard Deviation of Scaled Features: {}".format(np.std(X_scaled_wwq)))Mean of Scaled Features: 2.660809650821445e-15
Standard Deviation of Scaled Features: 1.0Centering and scaling in a pipeline
With regard to whether or not scaling is effective, the proof is in the pudding! We will See for ourselves whether or not scaling the features of the White Wine Quality dataset has any impact on its performance. We will use a k-NN classifier as part of a pipeline that includes scaling, and for the purposes of comparison
# Setup the pipeline steps: steps
steps_wwq = [('scaler', StandardScaler()),
('knn', KNeighborsClassifier())]
# Create the pipeline: pipeline
pipeline_wwq = Pipeline(steps_wwq)
# Create train and test sets
X_train_wwq, X_test_wwq, y_train_wwq, y_test_wwq = train_test_split(X_wwq,y_wwq, test_size=.3, random_state=42)
# Fit the pipeline to the training set: knn_scaled
knn_scaled_wwq = pipeline_wwq.fit(X_train_wwq, y_train_wwq)
# Instantiate and fit a k-NN classifier to the unscaled data
knn_unscaled_wwq = KNeighborsClassifier().fit(X_train_wwq, y_train_wwq)
# Compute and print metrics
print('Accuracy with Scaling: {}'.format(knn_scaled_wwq.score(X_test_wwq, y_test_wwq)))
print('Accuracy without Scaling: {}'.format(knn_unscaled_wwq.score(X_test_wwq, y_test_wwq)))Accuracy with Scaling: 0.7700680272108843
Accuracy without Scaling: 0.6979591836734694It looks like scaling has significantly improved model performance!
Bringing it all together I: Pipeline for classification
We will build a pipeline that includes scaling and hyperparameter tuning to classify wine quality.
X_wwq[:5]array([[7.000e+00, 2.700e-01, 3.600e-01, 2.070e+01, 4.500e-02, 4.500e+01,
1.700e+02, 1.001e+00, 3.000e+00, 4.500e-01, 8.800e+00],
[6.300e+00, 3.000e-01, 3.400e-01, 1.600e+00, 4.900e-02, 1.400e+01,
1.320e+02, 9.940e-01, 3.300e+00, 4.900e-01, 9.500e+00],
[8.100e+00, 2.800e-01, 4.000e-01, 6.900e+00, 5.000e-02, 3.000e+01,
9.700e+01, 9.951e-01, 3.260e+00, 4.400e-01, 1.010e+01],
[7.200e+00, 2.300e-01, 3.200e-01, 8.500e+00, 5.800e-02, 4.700e+01,
1.860e+02, 9.956e-01, 3.190e+00, 4.000e-01, 9.900e+00],
[7.200e+00, 2.300e-01, 3.200e-01, 8.500e+00, 5.800e-02, 4.700e+01,
1.860e+02, 9.956e-01, 3.190e+00, 4.000e-01, 9.900e+00]])# Setup the pipeline
steps_wwq = [('scaler', StandardScaler()),
('SVM', SVC())]
pipeline_wwq = Pipeline(steps_wwq)
# Specify the hyperparameter space
parameters_wwq = {'SVM__C':[1, 10, 100],
'SVM__gamma':[0.1, 0.01]}
# Create train and test sets
X_train_wwq, X_test_wwq, y_train_wwq, y_test_wwq = train_test_split(X_wwq,y_wwq, test_size=.2, random_state=21)
# Instantiate the GridSearchCV object: cv
cv_wwq = GridSearchCV(pipeline_wwq, param_grid=parameters_wwq, cv=3)
# Fit to the training set
cv_wwq.fit(X_train_wwq, y_train_wwq)
# Predict the labels of the test set: y_pred
y_pred_wwq = cv_wwq.predict(X_test_wwq)
# Compute and print metrics
print("Accuracy: {}".format(cv_wwq.score(X_test_wwq, y_test_wwq)))
print(classification_report(y_test_wwq, y_pred_wwq))
print("Tuned Model Parameters: {}".format(cv_wwq.best_params_))Accuracy: 0.7795918367346939
precision recall f1-score support
False 0.83 0.85 0.84 662
True 0.67 0.63 0.65 318
accuracy 0.78 980
macro avg 0.75 0.74 0.74 980
weighted avg 0.78 0.78 0.78 980
Tuned Model Parameters: {'SVM__C': 10, 'SVM__gamma': 0.1}Bringing it all together II: Pipeline for regression
We will return to the gapminder dataset, it had a lot of missing data. We will build a pipeline that imputes the missing data, scales the features, and fits an ElasticNet to the Gapminder data. We will then tune the l1_ratio of the ElasticNet using GridSearchCV.
X_gapminder = np.array([[3.48110590e+07, 2.73000000e+00, 1.00000000e-01, 3.32894466e+00,
2.45962000e+01, 1.23140000e+04, 1.29904900e+02, 2.95000000e+01],
[1.98422510e+07, 6.43000000e+00, 2.00000000e+00, 1.47435339e+00,
2.22508300e+01, 7.10300000e+03, 1.30124700e+02, 1.92000000e+02],
[4.03818600e+07, 2.24000000e+00, 5.00000000e-01, 4.78516998e+00,
2.75017000e+01, 1.46460000e+04, 1.18891500e+02, 1.54000000e+01],
[2.97502900e+06, 1.40000000e+00, 1.00000000e-01, 1.80410622e+00,
2.53554200e+01, 7.38300000e+03, 1.32810800e+02, 2.00000000e+01],
[2.13703480e+07, 1.96000000e+00, 1.00000000e-01, 1.80163133e+01,
2.75637300e+01, 4.13120000e+04, 1.17375500e+02, 5.20000000e+00],
[8.33146500e+06, 1.41000000e+00, 3.00000000e-01, 8.18316002e+00,
2.64674100e+01, 4.39520000e+04, 1.24139400e+02, 4.60000000e+00],
[8.86871300e+06, 1.99000000e+00, 1.00000000e-01, 5.10953829e+00,
2.56511700e+01, 1.43650000e+04, 1.28602400e+02, 4.33000000e+01],
[3.48587000e+05, 1.89000000e+00, 3.10000000e+00, 3.13192132e+00,
2.72459400e+01, 2.43730000e+04, 1.24386200e+02, 1.45000000e+01],
[1.48252473e+08, 2.38000000e+00, 6.00000000e-02, 3.19161002e-01,
2.03974200e+01, 2.26500000e+03, 1.25030700e+02, 5.59000000e+01],
[2.77315000e+05, 1.83000000e+00, 1.30000000e+00, 6.00827884e+00,
2.63843900e+01, 1.60750000e+04, 1.26394000e+02, 1.54000000e+01],
[9.52645300e+06, 1.42000000e+00, 2.00000000e-01, 6.48817388e+00,
2.61644300e+01, 1.44880000e+04, 1.29796800e+02, 7.20000000e+00],
[1.07791550e+07, 1.82000000e+00, 2.00000000e-01, 9.79733671e+00,
2.67591500e+01, 4.16410000e+04, 1.21822700e+02, 4.70000000e+00],
[3.06165000e+05, 2.91000000e+00, 2.40000000e+00, 1.36012592e+00,
2.70225500e+01, 8.29300000e+03, 1.20922400e+02, 2.01000000e+01],
[8.97352500e+06, 5.27000000e+00, 1.20000000e+00, 5.37539184e-01,
2.24183500e+01, 1.64600000e+03, 1.30272300e+02, 1.16300000e+02],
[6.94990000e+05, 2.51000000e+00, 2.00000000e-01, 6.01210310e-01,
2.28218000e+01, 5.66300000e+03, 1.25125800e+02, 4.81000000e+01],
[9.59991600e+06, 3.48000000e+00, 2.00000000e-01, 1.43182915e+00,
2.44333500e+01, 5.06600000e+03, 1.22415500e+02, 5.20000000e+01],
[1.96786600e+06, 2.86000000e+00, 2.49000000e+01, 2.54720549e+00,
2.21298400e+01, 1.38580000e+04, 1.33130700e+02, 6.38000000e+01],
[1.94769696e+08, 1.90000000e+00, 4.50000000e-01, 2.02377284e+00,
2.57862300e+01, 1.39060000e+04, 1.24874500e+02, 1.86000000e+01],
[7.51364600e+06, 1.43000000e+00, 1.00000000e-01, 6.69013908e+00,
2.65428600e+01, 1.53680000e+04, 1.28472100e+02, 1.37000000e+01],
[1.47090110e+07, 6.04000000e+00, 1.20000000e+00, 1.09419171e-01,
2.12715700e+01, 1.35800000e+03, 1.30665100e+02, 1.30400000e+02],
[8.82179500e+06, 6.48000000e+00, 3.50000000e+00, 3.13888013e-02,
2.15029100e+01, 7.23000000e+02, 1.34195500e+02, 1.08600000e+02],
[1.39336600e+07, 3.05000000e+00, 6.00000000e-01, 2.87547496e-01,
2.08049600e+01, 2.44200000e+03, 1.17552800e+02, 5.15000000e+01],
[1.95704180e+07, 5.17000000e+00, 5.30000000e+00, 2.95541639e-01,
2.36817300e+01, 2.57100000e+03, 1.27282300e+02, 1.13800000e+02],
[3.33632560e+07, 1.68000000e+00, 2.00000000e-01, 1.63503986e+01,
2.74521000e+01, 4.14680000e+04, 1.18057100e+02, 5.80000000e+00],
[1.11397400e+07, 6.81000000e+00, 3.40000000e+00, 4.78391264e-02,
2.14856900e+01, 1.75300000e+03, 1.27864000e+02, 1.68000000e+02],
[1.66459400e+07, 1.89000000e+00, 4.00000000e-01, 4.24025914e+00,
2.70154200e+01, 1.86980000e+04, 1.25541700e+02, 8.90000000e+00],
[4.49016600e+07, 2.43000000e+00, 5.00000000e-01, 1.47609182e+00,
2.49404100e+01, 1.04890000e+04, 1.24023500e+02, 1.97000000e+01],
[6.65414000e+05, 5.05000000e+00, 6.00000000e-02, 1.78853064e-01,
2.20613100e+01, 1.44000000e+03, 1.32135400e+02, 9.12000000e+01],
[3.83277100e+06, 5.10000000e+00, 3.50000000e+00, 3.84220477e-01,
2.18713400e+01, 5.02200000e+03, 1.31693500e+02, 7.26000000e+01],
[4.42950600e+06, 1.91000000e+00, 3.00000000e-01, 1.91193342e+00,
2.64789700e+01, 1.22190000e+04, 1.21350000e+02, 1.03000000e+01],
[1.92616470e+07, 4.91000000e+00, 3.70000000e+00, 3.61896603e-01,
2.25646900e+01, 2.85400000e+03, 1.31523700e+02, 1.16900000e+02],
[4.34415100e+06, 1.43000000e+00, 6.00000000e-02, 5.28790258e+00,
2.65962900e+01, 2.18730000e+04, 1.30392100e+02, 5.90000000e+00],
[1.12902390e+07, 1.50000000e+00, 1.00000000e-01, 2.70177717e+00,
2.50686700e+01, 1.77650000e+04, 1.26059400e+02, 6.30000000e+00],
[5.49530200e+06, 1.89000000e+00, 2.00000000e-01, 8.54150780e+00,
2.61328700e+01, 4.50170000e+04, 1.19581500e+02, 4.30000000e+00],
[8.09639000e+05, 3.76000000e+00, 2.60000000e+00, 6.12799524e-01,
2.33840300e+01, 2.50200000e+03, 1.29337600e+02, 8.10000000e+01],
[1.44476000e+07, 2.73000000e+00, 4.00000000e-01, 2.11051780e+00,
2.55884100e+01, 9.24400000e+03, 1.22986400e+02, 2.68000000e+01],
[7.89761220e+07, 2.95000000e+00, 6.00000000e-02, 2.51239420e+00,
2.67324300e+01, 9.97400000e+03, 1.25093100e+02, 3.14000000e+01],
[6.00419900e+06, 2.32000000e+00, 8.00000000e-01, 1.06776463e+00,
2.63675100e+01, 7.45000000e+03, 1.19932100e+02, 2.16000000e+01],
[6.86223000e+05, 5.31000000e+00, 4.70000000e+00, 6.79825323e+00,
2.37664000e+01, 4.01430000e+04, 1.32039200e+02, 1.18400000e+02],
[4.50063800e+06, 5.16000000e+00, 8.00000000e-01, 8.37456442e-02,
2.08850900e+01, 1.08800000e+03, 1.25794800e+02, 6.04000000e+01],
[1.33994100e+06, 1.62000000e+00, 1.20000000e+00, 1.30313789e+01,
2.62644600e+01, 2.47430000e+04, 1.29516100e+02, 5.50000000e+00],
[8.43206000e+05, 2.74000000e+00, 1.00000000e-01, 1.27777956e+00,
2.65307800e+01, 7.12900000e+03, 1.27476800e+02, 2.40000000e+01],
[5.31417000e+06, 1.85000000e+00, 1.00000000e-01, 1.06441143e+01,
2.67333900e+01, 4.21220000e+04, 1.26564500e+02, 3.30000000e+00],
[6.23095290e+07, 1.97000000e+00, 4.00000000e-01, 5.99902073e+00,
2.58532900e+01, 3.75050000e+04, 1.20014600e+02, 4.30000000e+00],
[1.47374100e+06, 4.28000000e+00, 5.30000000e+00, 1.07953932e+00,
2.40762000e+01, 1.58000000e+04, 1.30362500e+02, 6.80000000e+01],
[1.58674900e+06, 5.80000000e+00, 1.70000000e+00, 2.51002328e-01,
2.16502900e+01, 1.56600000e+03, 1.30208000e+02, 8.74000000e+01],
[4.34329000e+06, 1.79000000e+00, 1.00000000e-01, 1.41942978e+00,
2.55494200e+01, 5.90000000e+03, 1.30578900e+02, 1.93000000e+01],
[8.06659060e+07, 1.37000000e+00, 1.00000000e-01, 9.49724676e+00,
2.71650900e+01, 4.11990000e+04, 1.24904400e+02, 4.40000000e+00],
[2.31159190e+07, 4.19000000e+00, 1.80000000e+00, 3.66600849e-01,
2.28424700e+01, 2.90700000e+03, 1.28295300e+02, 7.99000000e+01],
[1.11617550e+07, 1.46000000e+00, 1.00000000e-01, 8.66123553e+00,
2.63378600e+01, 3.21970000e+04, 1.22934200e+02, 4.90000000e+00],
[1.41066870e+07, 4.12000000e+00, 8.00000000e-01, 8.35594820e-01,
2.52994700e+01, 6.96000000e+03, 1.20959600e+02, 3.69000000e+01],
[1.04273560e+07, 5.34000000e+00, 1.40000000e+00, 1.26964400e-01,
2.25244900e+01, 1.23000000e+03, 1.32276500e+02, 1.21000000e+02],
[1.56129300e+06, 5.25000000e+00, 2.50000000e+00, 1.56376432e-01,
2.16433800e+01, 1.32600000e+03, 1.30762700e+02, 1.27600000e+02],
[7.48096000e+05, 2.74000000e+00, 1.20000000e+00, 2.07341531e+00,
2.36846500e+01, 5.20800000e+03, 1.25151200e+02, 4.19000000e+01],
[9.70513000e+06, 3.50000000e+00, 2.00000000e+00, 2.49306755e-01,
2.36630200e+01, 1.60000000e+03, 1.25346100e+02, 8.33000000e+01],
[7.25947000e+06, 3.27000000e+00, 8.00000000e-01, 1.18745352e+00,
2.51087200e+01, 4.39100000e+03, 1.22962100e+02, 2.65000000e+01],
[1.00506990e+07, 1.33000000e+00, 6.00000000e-02, 5.45323172e+00,
2.71156800e+01, 2.33340000e+04, 1.28696800e+02, 7.20000000e+00],
[3.10033000e+05, 2.12000000e+00, 3.00000000e-01, 6.82190305e+00,
2.72068700e+01, 4.22940000e+04, 1.18738100e+02, 2.70000000e+00],
[1.19707011e+09, 2.64000000e+00, 3.20000000e-01, 1.52084942e+00,
2.09595600e+01, 3.90100000e+03, 1.23127400e+02, 6.56000000e+01],
[2.35360765e+08, 2.48000000e+00, 2.00000000e-01, 1.75504422e+00,
2.18557600e+01, 7.85600000e+03, 1.26421600e+02, 3.62000000e+01],
[7.25306930e+07, 1.88000000e+00, 2.00000000e-01, 7.89221094e+00,
2.53100300e+01, 1.59550000e+04, 1.25185900e+02, 2.14000000e+01],
[4.48014500e+06, 2.00000000e+00, 2.00000000e-01, 9.88253103e+00,
2.76532500e+01, 4.77130000e+04, 1.24780100e+02, 4.50000000e+00],
[7.09380800e+06, 2.92000000e+00, 2.00000000e-01, 1.00011881e+01,
2.71315100e+01, 2.85620000e+04, 1.21083800e+02, 4.90000000e+00],
[5.93192340e+07, 1.39000000e+00, 3.00000000e-01, 7.46594241e+00,
2.64802000e+01, 3.74750000e+04, 1.23703000e+02, 4.10000000e+00],
[2.71734400e+06, 2.39000000e+00, 1.70000000e+00, 4.39145647e+00,
2.40042100e+01, 8.95100000e+03, 1.25368500e+02, 1.89000000e+01],
[1.27317900e+08, 1.34000000e+00, 6.00000000e-02, 9.53660569e+00,
2.35000400e+01, 3.48000000e+04, 1.21965100e+02, 3.40000000e+00],
[1.59159660e+07, 2.51000000e+00, 1.00000000e-01, 1.47181043e+01,
2.62907800e+01, 1.87970000e+04, 1.28851700e+02, 2.59000000e+01],
[3.82444420e+07, 4.76000000e+00, 6.30000000e+00, 2.66308378e-01,
2.15925800e+01, 2.35800000e+03, 1.29934100e+02, 7.10000000e+01],
[2.14421500e+06, 1.50000000e+00, 6.00000000e-01, 3.34184866e+00,
2.64569300e+01, 2.09770000e+04, 1.29574600e+02, 1.05000000e+01],
[4.10938900e+06, 1.57000000e+00, 1.00000000e-01, 3.99672180e+00,
2.72011700e+01, 1.41580000e+04, 1.27503700e+02, 1.13000000e+01],
[1.97219400e+06, 3.34000000e+00, 2.36000000e+01, 8.61766942e-03,
2.19015700e+01, 2.04100000e+03, 1.31136100e+02, 1.14200000e+02],
[3.67278200e+06, 5.19000000e+00, 1.60000000e+00, 1.57352183e-01,
2.18953700e+01, 5.88000000e+02, 1.31255500e+02, 1.00900000e+02],
[3.21980200e+06, 1.42000000e+00, 1.00000000e-01, 4.49848339e+00,
2.68610200e+01, 2.32230000e+04, 1.30822600e+02, 8.20000000e+00],
[4.85079000e+05, 1.63000000e+00, 3.00000000e-01, 2.21680797e+01,
2.74340400e+01, 9.50010000e+04, 1.22370500e+02, 2.80000000e+00],
[1.99267980e+07, 4.79000000e+00, 2.00000000e-01, 9.94221476e-02,
2.14034700e+01, 1.52800000e+03, 1.32837100e+02, 6.67000000e+01],
[1.39046710e+07, 5.78000000e+00, 1.12000000e+01, 8.24698808e-02,
2.20346800e+01, 6.74000000e+02, 1.33939000e+02, 1.01100000e+02],
[2.71974190e+07, 2.05000000e+00, 5.00000000e-01, 7.75223395e+00,
2.47306900e+01, 1.99680000e+04, 1.23859300e+02, 8.00000000e+00],
[3.21026000e+05, 2.38000000e+00, 6.00000000e-02, 3.27772577e+00,
2.32199100e+01, 1.20290000e+04, 1.23322300e+02, 1.60000000e+01],
[1.42234030e+07, 6.82000000e+00, 1.00000000e+00, 4.10788666e-02,
2.17888100e+01, 1.60200000e+03, 1.28030800e+02, 1.48300000e+02],
[4.06392000e+05, 1.38000000e+00, 1.00000000e-01, 6.18277102e+00,
2.76836100e+01, 2.78720000e+04, 1.24157100e+02, 6.60000000e+00],
[3.41455200e+06, 4.94000000e+00, 7.00000000e-01, 6.13103977e-01,
2.26229500e+01, 3.35600000e+03, 1.29987500e+02, 1.03000000e+02],
[1.23801300e+06, 1.58000000e+00, 9.00000000e-01, 3.07876290e+00,
2.51566900e+01, 1.46150000e+04, 1.30878600e+02, 1.58000000e+01],
[1.14972821e+08, 2.35000000e+00, 3.00000000e-01, 4.26117187e+00,
2.74246800e+01, 1.58260000e+04, 1.22121600e+02, 1.79000000e+01],
[4.11116800e+06, 1.49000000e+00, 4.00000000e-01, 1.31332119e+00,
2.42369000e+01, 3.89000000e+03, 1.29942400e+02, 1.76000000e+01],
[2.62966600e+06, 2.37000000e+00, 6.00000000e-02, 3.75948682e+00,
2.48838500e+01, 7.56300000e+03, 1.29750400e+02, 3.48000000e+01],
[3.13505440e+07, 2.44000000e+00, 1.00000000e-01, 1.59408314e+00,
2.56318200e+01, 6.09100000e+03, 1.26528400e+02, 3.58000000e+01],
[2.29948670e+07, 5.54000000e+00, 1.14000000e+01, 1.04748301e-01,
2.19353600e+01, 8.64000000e+02, 1.35394900e+02, 1.14400000e+02],
[5.10300060e+07, 2.05000000e+00, 6.00000000e-01, 1.91053400e-01,
2.14493200e+01, 2.89100000e+03, 1.23142100e+02, 8.72000000e+01],
[2.63251830e+07, 2.90000000e+00, 4.00000000e-01, 1.05412983e-01,
2.07634400e+01, 1.86600000e+03, 1.25556100e+02, 5.07000000e+01],
[1.65198620e+07, 1.77000000e+00, 2.00000000e-01, 1.05330281e+01,
2.60154100e+01, 4.73880000e+04, 1.21695000e+02, 4.80000000e+00],
[4.28538000e+06, 2.12000000e+00, 1.00000000e-01, 8.00908440e+00,
2.77689300e+01, 3.21220000e+04, 1.18742100e+02, 6.40000000e+00],
[5.59452400e+06, 2.72000000e+00, 2.00000000e-01, 7.78151613e-01,
2.57729100e+01, 4.06000000e+03, 1.23479200e+02, 2.81000000e+01],
[1.50851300e+07, 7.59000000e+00, 8.00000000e-01, 6.34371088e-02,
2.12195800e+01, 8.43000000e+02, 1.35102100e+02, 1.41300000e+02],
[1.51115683e+08, 6.02000000e+00, 3.60000000e+00, 6.14689662e-01,
2.30332200e+01, 4.68400000e+03, 1.35492000e+02, 1.40900000e+02],
[4.77163300e+06, 1.96000000e+00, 1.00000000e-01, 1.05297688e+01,
2.69342400e+01, 6.52160000e+04, 1.26026600e+02, 3.60000000e+00],
[2.65228100e+06, 2.89000000e+00, 1.00000000e-01, 1.55720805e+01,
2.62410900e+01, 4.77990000e+04, 1.26887000e+02, 1.19000000e+01],
[1.63096985e+08, 3.58000000e+00, 1.00000000e-01, 9.35618056e-01,
2.22991400e+01, 4.18700000e+03, 1.26519600e+02, 9.55000000e+01],
[3.49867900e+06, 2.61000000e+00, 9.00000000e-01, 2.22379634e+00,
2.62695900e+01, 1.40330000e+04, 1.22682900e+02, 2.10000000e+01],
[6.54026700e+06, 4.07000000e+00, 9.00000000e-01, 5.30746337e-01,
2.50150600e+01, 1.98200000e+03, 1.20052400e+02, 6.97000000e+01],
[6.04713100e+06, 3.06000000e+00, 3.00000000e-01, 6.98581746e-01,
2.55422300e+01, 6.68400000e+03, 1.23615000e+02, 2.57000000e+01],
[2.86420480e+07, 2.58000000e+00, 4.00000000e-01, 1.45013444e+00,
2.47704100e+01, 9.24900000e+03, 1.19636800e+02, 2.32000000e+01],
[9.02971150e+07, 3.26000000e+00, 6.00000000e-02, 8.42120697e-01,
2.28726300e+01, 5.33200000e+03, 1.22345900e+02, 3.34000000e+01],
[3.85257520e+07, 1.33000000e+00, 1.00000000e-01, 8.27076715e+00,
2.66738000e+01, 1.99960000e+04, 1.29676500e+02, 6.70000000e+00],
[1.05774580e+07, 1.36000000e+00, 5.00000000e-01, 5.48692640e+00,
2.66844500e+01, 2.77470000e+04, 1.27263100e+02, 4.10000000e+00],
[1.38896200e+06, 2.20000000e+00, 6.00000000e-02, 4.87020615e+01,
2.81313800e+01, 1.26076000e+05, 1.26315300e+02, 9.50000000e+00],
[2.07416690e+07, 1.34000000e+00, 1.00000000e-01, 4.38344907e+00,
2.54106900e+01, 1.80320000e+04, 1.28755300e+02, 1.61000000e+01],
[1.43123163e+08, 1.49000000e+00, 1.00000000e+00, 1.19827176e+01,
2.60113100e+01, 2.25060000e+04, 1.28490300e+02, 1.35000000e+01],
[9.75031400e+06, 5.06000000e+00, 2.90000000e+00, 5.42444698e-02,
2.25545300e+01, 1.17300000e+03, 1.35100500e+02, 7.83000000e+01],
[1.22297030e+07, 5.11000000e+00, 8.00000000e-01, 4.61633711e-01,
2.19274300e+01, 2.16200000e+03, 1.30279500e+02, 7.58000000e+01],
[9.10953500e+06, 1.41000000e+00, 1.00000000e-01, 5.27122268e+00,
2.65149500e+01, 1.25220000e+04, 1.30375500e+02, 8.00000000e+00],
[5.52183800e+06, 5.13000000e+00, 1.60000000e+00, 1.18255775e-01,
2.25313900e+01, 1.28900000e+03, 1.34716000e+02, 1.79100000e+02],
[4.84964100e+06, 1.28000000e+00, 1.00000000e-01, 4.11444075e+00,
2.38399600e+01, 6.59910000e+04, 1.21173600e+02, 2.80000000e+00],
[5.39671000e+06, 1.31000000e+00, 6.00000000e-02, 6.90165446e+00,
2.69271700e+01, 2.46700000e+04, 1.29528000e+02, 8.80000000e+00],
[2.03059900e+06, 1.43000000e+00, 6.00000000e-02, 8.51182820e+00,
2.74398300e+01, 3.08160000e+04, 1.29923100e+02, 3.70000000e+00],
[9.13258900e+06, 7.06000000e+00, 6.00000000e-01, 6.82188892e-02,
2.19691700e+01, 6.15000000e+02, 1.31531800e+02, 1.68500000e+02],
[5.03488110e+07, 2.54000000e+00, 1.79000000e+01, 9.42796037e+00,
2.68553800e+01, 1.22630000e+04, 1.30994900e+02, 6.61000000e+01],
[4.58170160e+07, 1.42000000e+00, 4.00000000e-01, 7.29308876e+00,
2.74997500e+01, 3.46760000e+04, 1.22045300e+02, 5.00000000e+00],
[1.99495530e+07, 2.32000000e+00, 6.00000000e-02, 5.80791088e-01,
2.19667100e+01, 6.90700000e+03, 1.24861500e+02, 1.17000000e+01],
[3.44701380e+07, 4.79000000e+00, 1.00000000e+00, 3.82117945e-01,
2.24048400e+01, 3.24600000e+03, 1.29719900e+02, 8.47000000e+01],
[5.06657000e+05, 2.41000000e+00, 1.00000000e+00, 4.74113997e+00,
2.54988700e+01, 1.34700000e+04, 1.24635800e+02, 2.64000000e+01],
[1.15375000e+06, 3.70000000e+00, 2.59000000e+01, 9.49860795e-01,
2.31696900e+01, 5.88700000e+03, 1.31879300e+02, 1.12200000e+02],
[9.22633300e+06, 1.92000000e+00, 1.00000000e-01, 5.31568840e+00,
2.63762900e+01, 4.34210000e+04, 1.22947300e+02, 3.20000000e+00],
[7.64654200e+06, 1.47000000e+00, 4.00000000e-01, 5.33305762e+00,
2.62019500e+01, 5.50200000e+04, 1.19646500e+02, 4.70000000e+00],
[7.25407200e+06, 3.70000000e+00, 2.00000000e-01, 4.53167660e-01,
2.37796600e+01, 2.00100000e+03, 1.29965700e+02, 5.62000000e+01],
[4.28447440e+07, 5.54000000e+00, 5.80000000e+00, 1.54672995e-01,
2.24779200e+01, 2.03000000e+03, 1.30832800e+02, 7.24000000e+01],
[6.64532550e+07, 1.48000000e+00, 1.30000000e+00, 3.83510189e+00,
2.30080300e+01, 1.22160000e+04, 1.20496900e+02, 1.56000000e+01],
[6.05293700e+06, 4.88000000e+00, 3.20000000e+00, 2.51983337e-01,
2.18787500e+01, 1.21900000e+03, 1.31024800e+02, 9.64000000e+01],
[1.31537200e+06, 1.80000000e+00, 1.50000000e+00, 3.19577173e+01,
2.63966900e+01, 3.08750000e+04, 1.24993900e+02, 2.49000000e+01],
[1.04080910e+07, 2.04000000e+00, 6.00000000e-02, 2.44066948e+00,
2.51569900e+01, 9.93800000e+03, 1.28629100e+02, 1.94000000e+01],
[7.03443570e+07, 2.15000000e+00, 6.00000000e-02, 4.02190259e+00,
2.67037100e+01, 1.64540000e+04, 1.24067500e+02, 2.22000000e+01],
[3.10144270e+07, 6.34000000e+00, 6.40000000e+00, 1.00852839e-01,
2.23583300e+01, 1.43700000e+03, 1.34520400e+02, 8.93000000e+01],
[4.60284760e+07, 1.38000000e+00, 1.10000000e+00, 7.03235908e+00,
2.54237900e+01, 8.76200000e+03, 1.31496200e+02, 1.29000000e+01],
[6.16896200e+07, 1.87000000e+00, 2.00000000e-01, 8.52646682e+00,
2.73924900e+01, 3.77390000e+04, 1.24084500e+02, 5.60000000e+00],
[3.04473143e+08, 2.07000000e+00, 6.00000000e-01, 1.85459917e+01,
2.84569800e+01, 5.03840000e+04, 1.18477700e+02, 7.70000000e+00],
[3.35083200e+06, 2.11000000e+00, 5.00000000e-01, 2.48976355e+00,
2.63912300e+01, 1.53170000e+04, 1.24260400e+02, 1.30000000e+01],
[2.69527190e+07, 2.46000000e+00, 1.00000000e-01, 4.47666902e+00,
2.53205400e+01, 3.73300000e+03, 1.24346200e+02, 4.92000000e+01],
[8.65893420e+07, 1.86000000e+00, 4.00000000e-01, 1.47934658e+00,
2.09163000e+01, 4.08500000e+03, 1.21936700e+02, 2.62000000e+01],
[1.31145790e+07, 5.88000000e+00, 1.36000000e+01, 1.48981514e-01,
2.06832100e+01, 3.03900000e+03, 1.32449300e+02, 9.49000000e+01],
[1.34954620e+07, 3.85000000e+00, 1.51000000e+01, 6.54323190e-01,
2.20266000e+01, 1.28600000e+03, 1.31974500e+02, 9.83000000e+01]])X_gapminder = pd.read_csv("datasets/X_gapminder.csv").values
X_gapminder[:5]array([[3.48110590e+07, 2.73000000e+00, 1.00000000e-01, 3.32894466e+00,
2.45962000e+01, 1.23140000e+04, 1.29904900e+02, 2.95000000e+01],
[1.98422510e+07, 6.43000000e+00, 2.00000000e+00, 1.47435339e+00,
2.22508300e+01, 7.10300000e+03, 1.30124700e+02, 1.92000000e+02],
[4.03818600e+07, 2.24000000e+00, 5.00000000e-01, 4.78516998e+00,
2.75017000e+01, 1.46460000e+04, 1.18891500e+02, 1.54000000e+01],
[2.97502900e+06, 1.40000000e+00, 1.00000000e-01, 1.80410622e+00,
2.53554200e+01, 7.38300000e+03, 1.32810800e+02, 2.00000000e+01],
[2.13703480e+07, 1.96000000e+00, 1.00000000e-01, 1.80163133e+01,
2.75637300e+01, 4.13120000e+04, 1.17375500e+02, 5.20000000e+00]])y_gapminder = np.array([75.3, 58.3, 75.5, 72.5, 81.5, 80.4, 70.6, 72.2, 68.4, 75.3, 70.1,
79.4, 70.7, 63.2, 67.6, 70.9, 61.2, 73.9, 73.2, 59.4, 57.4, 66.2,
56.6, 80.7, 54.8, 78.9, 75.1, 62.6, 58.6, 79.7, 55.9, 76.5, 77.8,
78.7, 61. , 74. , 70.1, 74.1, 56.7, 60.4, 74. , 65.7, 79.4, 81. ,
57.5, 62.2, 72.1, 80. , 62.7, 79.5, 70.8, 58.3, 51.3, 63. , 61.7,
70.9, 73.8, 82. , 64.4, 69.5, 76.9, 79.4, 80.9, 81.4, 75.5, 82.6,
66.1, 61.5, 72.3, 77.6, 45.2, 61. , 72. , 80.7, 63.4, 51.4, 74.5,
78.2, 55.8, 81.4, 63.6, 72.1, 75.7, 69.6, 63.2, 73.3, 55. , 60.8,
68.6, 80.3, 80.2, 75.2, 59.7, 58. , 80.7, 74.6, 64.1, 77.1, 58.2,
73.6, 76.8, 69.4, 75.3, 79.2, 80.4, 73.4, 67.6, 62.2, 64.3, 76.4,
55.9, 80.9, 74.8, 78.5, 56.7, 55. , 81.1, 74.3, 67.4, 69.1, 46.1,
81.1, 81.9, 69.5, 59.7, 74.1, 60. , 71.3, 76.5, 75.1, 57.2, 68.2,
79.5, 78.2, 76. , 68.7, 75.4, 52. , 49. ])y_gapminder = pd.read_csv("datasets/y_gapminder.csv").values
y_gapminder[:5]array([[75.3],
[58.3],
[75.5],
[72.5],
[81.5]])# Setup the pipeline steps: steps
steps_gapminder = [('imputation', SimpleImputer(missing_values=np.nan, strategy="mean")),
("scaler", StandardScaler()),
("elasticnet", ElasticNet())]
# Create the pipeline: pipeline
pipeline_gapminder = Pipeline(steps_gapminder)
# Specify the hyperparameter space
parameters_gapminder = {"elasticnet__l1_ratio":np.linspace(0,1,30)}
# Create train and test sets
X_train_gapminder, X_test_gapminder, y_train_gapminder, y_test_gapminder = train_test_split(X_gapminder,
y_gapminder,
test_size=.4,
random_state=42)
# Create the GridSearchCV object: gm_cv
gm_cv_gapminder = GridSearchCV(pipeline_gapminder, param_grid=parameters_gapminder, cv=3)
# Fit to the training set
gm_cv_gapminder.fit(X_train_gapminder, y_train_gapminder)
# Compute and print the metrics
r2_gapminder = gm_cv_gapminder.score(X_test_gapminder, y_test_gapminder)
print("Tuned ElasticNet Alpha: {}".format(gm_cv_gapminder.best_params_))
print("Tuned ElasticNet R squared: {}".format(r2_gapminder))Tuned ElasticNet Alpha: {'elasticnet__l1_ratio': 1.0}
Tuned ElasticNet R squared: 0.8862016570888216